Another journey into the microscopic world of the various niches provided by fish holobionts with an emphasis on the skin mucosal communities which are the fish’s first line of defence (Banner, personal communication). The skin microbiota of seabass (Dicentrarchus labrax) fingerlings was evaluated for microbial composition, function, and dysbiotic variation during natural outbreaks of Photobacterium damselae piscicida and Vibrio harveyi, while microbial profiling continued throughout mitigation with the broad-spectrum gram negative fluoroquinolone antibiotic, Flumequine. Meta-taxonomy assayed perturbations in prokaryotic communities whilst the antibiotic persisted in muscle and skin from four to 14 days. Fish were reared in ponds fed by water from the Alvor Estuary in Southern Portugal for six months until weighing an average of 57 g where the concluding samples were collected at seven months when the fish weighed ~70 g (Rosado et al. 2022).
Disease and antibiotic treatments progressively raised skin microbial diversity and evenness compared to healthy controls, where diversity declination was associated with recovery. 49 prokaryotic phyla and 926 genera were detected, where six phyla and 32 genera were enriched in healthy and recuperating fish. Variations in community composition were associated with the onset of disease and later during antibiotic treatment. The profusion of Photobacterium damselae piscicida was comparable in the skin communities where it was elevated in diseased fish and gradually diminished during antibiotic treatment and recovery. In contrast Vibrio species abundance increased with the onset of disease and was constrained by antibiotics yet re-emerged during recouperation (Fig 1.).
Remarkably from 74 core affiliate amplicon sequence variants (ASVs) found in all four groups, six, 20, 30, and 67 were detected in samples from healthy, diseased, treated, and recovered seabass respectively. Two ASVs were unique to diseased fingerlings and two others were only found in treated fish, while 38 were common to those that recovered. One of the unique phylotypes from diseased fish was a Photobacterium species while a further two were likely pathogenic Vibrio. Two and four other putative commensal Vibrio ASVs were core members of the skin communities of recovering fish. The skin microbiome profiles of healthy fingerlings were dissimilar to diseased or recovering seabass which were more alike (Fig 2.; Rosado et al. 2022).
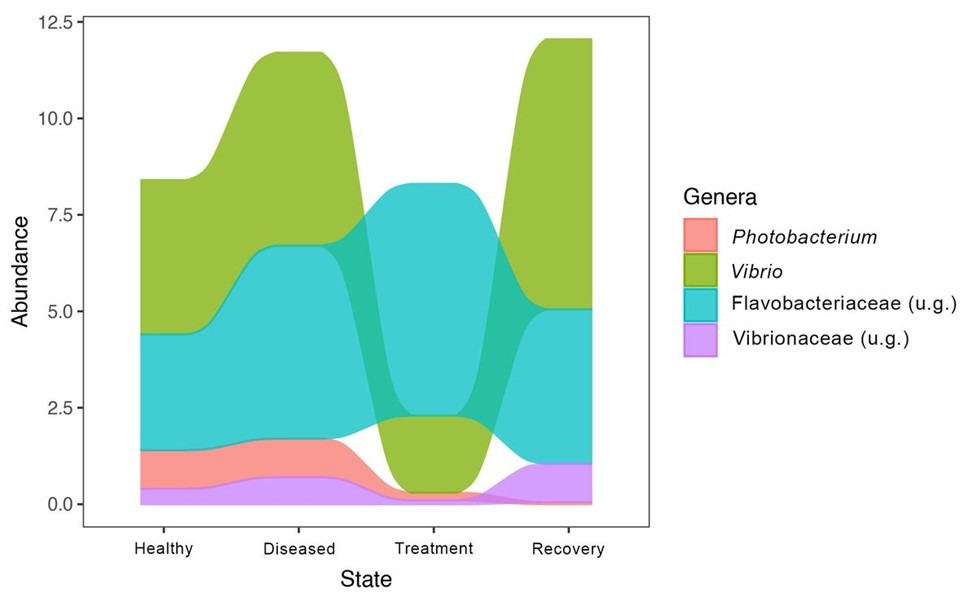
Fig 1. The relative abundances of the predominant microbial genera and families in the skin mucosae of healthy, diseased, treated, and recovering Dicentrarchus labrax fingerlings. Image courtesy of Rosado et al. 2022 and the Creative Commons Attribution Licence 4.0. https://creativecommons.org/licenses/by/4.0/
Kyoto encyclopaedia of genes and genomes (KEGG) pathway metadata and linear discriminant analysis effect size (LEfSe) discerned the metabolome of seabass skin microbiota where three, five, and seven metabolites were elevated in the healthy, treated, and recovered groups but nil were enriched in the skin of diseased fish. 67 and 57 percent of metabolic pathways were associated with biosynthesis in healthy and recovering skin biota, whereas 33, 100, and 43 percent were related to biochemical utilisation/assimilation in the skin of healthy, treated, and recovering fish respectively. Pathways associated with amino acid degradation and fatty acid and lipid biosynthesis were upregulated in the metabolome of healthy fish, while the micro-consortia of treated fish skins exhibited carbohydrate and secondary metabolite-degradative competencies. Nonetheless carbohydrate digestion and vitamin, fatty acid, and purine nucleotide-biosynthesis monopolised the skin metabolome of recovering fingerlings (Fig 3.; Rosado et al. 2022).
In contrast to the findings of extensive microbiome research, microbial diversity increased during the onset of disease (Rosado et al. 2022). Dysbiotic purging typically provides a vacant niche for inhabitation with environmental microbes and the development of a pathobiome yet instead there was an upsurge of putative opportunistic and aetiological taxa like Aureispira species and members of the families Flavobacteriaceae and Vibrionaceae. However, a perturbation of key microbial phylotypes was observed which epitomises dysbiosis (Rosado et al. 2022).
A single strain of Photobacterium damselae was represented in the skin microbiome of all diseased and undiseased fish where a previous study had highlighted the pathogen’s ability to elicit dysbiosis of seabass skin while its prevalence remained stable or increased (Rosado et al. 2019a; Rosado et al. 2022) and the skin and gill microbiomes of yellowtail kingfish experienced symbiotic dissociation during the onset of enteritis (Legrand et al. 2018).
Significant taxonomic shifts in microbial community composition were observed in 77 percent of diseased fish, where changes in the mucosal biochemical profile may have nurtured select lineages at the expense of others. Yet remarkably, the metabolome remained unaltered. The upsurge in diversity coincided with the recruitment of bacteria with analogous metabolic functions and donations (Kelly & Salinas 2017). Hence these affiliates offered an operational rather than a dysfunctional pathobiome (Rosado et al. 2022) perhaps because it is never in a pathogen’s interest to kill its host.
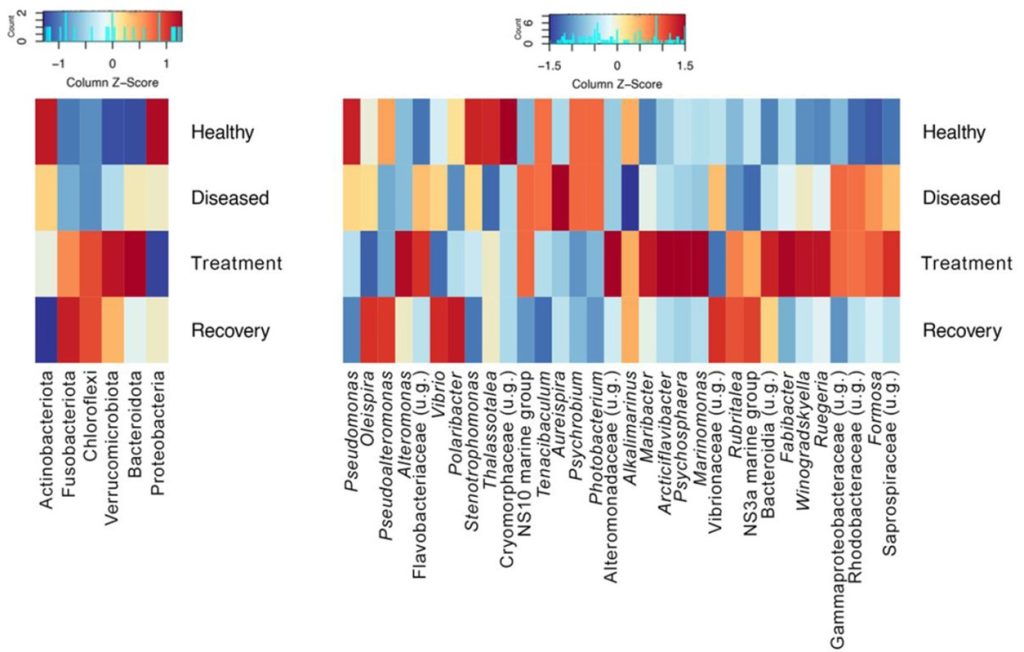
Fig 2. Heatmaps of the relative abundances of the prokaryotic phyla and amplicon sequence variants (ASVs; phylotypes) in the skin microbiomes of healthy, diseased, treated, and recovering Dicentrarchus labrax fingerlings. Image courtesy of Rosado et al. 2022 and the Creative Commons Attribution Licence 4.0. https://creativecommons.org/licenses/by/4.0/
An escalation of skin mucosal microbiome diversity, disease amelioration, and a downturn in pathogenic strains was observed on the eighth day of flumequine administration (Rosado et al. 2022). Nevertheless, there was a concurrent increase of Flavobacteriaceae many of which are pathogenic (Austin & Austin 2016, cited in Rosado et al. 2022) and Alteromonas species known to liberate Photobacterium damselae and Vibrio species antimicrobials (Stevens et al. 2016) and withstand amoxicillin, erythromycin, and gentamicin (Wu et al. 2019, cited in Rosado et al. 2022). Antibiotics have increased microbial diversity but it is not the norm (López Nadal et al. 2018; Rosado et al. 2019a) but in most instances dysbiosis and pathogenicity were not pre-existing which proposes antibiotics, disease, and dysbiotic purging act synergistically to increase diversity (Rosado et al. 2022).
Significant skin metabolome disparities escalated the breakdown of complex carbohydrates (glycans) and secondary metabolites throughout antibiotic administration (Rosado et al. 2022) where the former may be shielding (Hooper & Gordon 2001; Kelly & Salinas 2017) insofar as carbohydrates assist host cell, commensal, and pathogen cell-to-cell or mucus-to-cell adhesions (Hooper & Gordon 2001; Spor et al. 2011). Hence digestion of this kind may prove detrimental (Rosado et al. 2022) whereas streptomycin remains an immunosuppressive (Colorni et al. 1998).
Putative pathogenic Vibrio species were enriched in the post-antibiotic skin microbiome of recovered fingerlings compared with non-medicated undiseased controls, which may be indicative of partial convalescence (Rosado et al. 2022) because recuperated coral mucosal microbiomes revert to their prestressed profiles (Santoro et al. 2020). Notwithstanding, commonality was observed amongst the community compositions of healthy and recovered fish (Rosado et al. 2022).
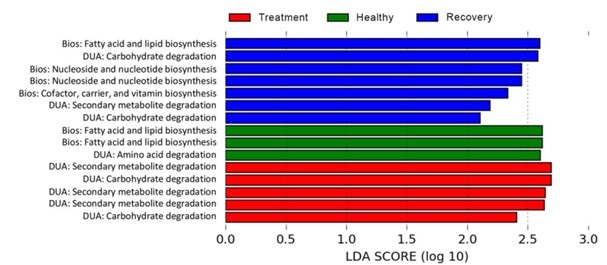
Fig 3. Skin metabolome analysis of healthy, antibiotic-treated, and recuperating Dicentrarchus labrax fingerlings: biosynthesis [Bios]; degradation/utilization/assimilation [DUA]. Image courtesy of Rosado et al. 2022 and the Creative Commons Attribution Licence 4.0. https://creativecommons.org/licenses/by/4.0/
Next we conclude this ambitious journey with an exploration of the gut and gill microbiota of saltwater fish and their dietary, prebiotic, and probiotic manipulations.
References
Adams, J. (2018) Interesting Ecology Shift of Blacktail and Orangeface Butterflyfish. ReefBuilders.com. https://reefbuilders.com/2018/12/18/blacktail-and-redface-butterflyfish/
Akhter, N., Wu, B., Memon, A., M. & Mohsin, M. (2015) Probiotics and prebiotics associated with aquaculture: a review. Fish Shellfish Immunol. 45, 733-741. https://www.doi.org/10.1016/j.fsi.2015.05.038
Amann, R., I., Ludwig, W. & Schleifer, K., H. (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 59, 143-169.
Andreoni, F. & Magnani, M. (2014) Photobacteriosis: Prevention and Diagnosis. Journal of immunology research. 2014,.
Aslett, C., G. (2023a) Coral Immunity Part I. https://www.reefranch.co.uk/
Aslett, C., G. (2023b) Holosystemics Part IV: Dysbiosis and the Microscopic Coral Alliance. https://www.reefranch.co.uk/
Aslett, C., G. (2023c) Fish Immunostimulation. https://www.reefranch.co.uk/
Aslett, C., G. (2024) The Complete Reef Aquarist, for hobbyists, the trade and academics – A Conservation Manual. Reef Ranch Publishing Ltd, Leeds, West Yorkshire, UK.
Austin, B. & Austin, D., A. (2016) Bacterial fish pathogens, 6th ed. Springer, Stirling, UK
Austin, B. (1982) Taxonomy of bacteria isolated from a coastal, marine fish-rearing unit. J. Appl. Bacteriol. 53, 253-268. https://www.doi.org/10.1111/j.1365-2672.1982.tb04684.x
Banner, C., Oregon Department of Fish & Wildlife.
Bass, D., Stentiford. G., D., Wang, H-C., Koskella, B. & Tyler, C., R. (2019) The pathobiome in animal and plant diseases. Trends Ecol Evol. 34(11), 996-1008.
Bell, G., R., Hoskins, G., E. & Hodgkiss, W. (1971) Aspects of the characterization, identification, and ecology of the bacterial flora associated with the surface of stream-incubating Pacific salmon (Oncorhynchus) eggs. J. Fish. Board Can. 28, 1511-1525. https://www.doi.org/10.1139/f71-232
Birrell, C., Mccook, L., Willis, B. & Diaz-Pulido, G. (2008) Effects of benthic algae on the replenishment of corals and the implications for the resilience of coral reefs. Oceanography and marine biology. 46,.
Blazer, V., Phillips, S. & Pendleton, E. (2016) Fish Health, Fungal Infections, and Pfiesteria: The Role of the U.S. Geological Survey. U.S. Geological Survey Fact Sheet. pp 114-198.
Boilard, A., Dubé, C., E., Gruet, C., Mercière, A., Hernandez-Agreda, A. & Derome, N. (2020) Defining Coral Bleaching as a Microbial Dysbiosis within the Coral Holobiont. Microorganisms. 8, 1682. https://doi.org/10.3390/microorganisms8111682
Bone, Q., Marshall, N., B. & Blaxter, J., H., S. (1995) Biology of Fishes. Glasgow: Blackie Academic & Professional. https://www.doi.org/10.1007/978-1-4615-2664-3
Bozzi, D., Rasmussen, J., A., Carøe, C., et al. (2021) Salmon gut microbiota correlates with disease infection status: potential for monitoring health in farmed animals. Anim Microbiome. 3(1), 1-17.
Brandley, B., K. & Schnaar, R., L. (1986) Cell-Surface Carbohydrates in Cell Recognition and Response. J Leukoc Biol 40, 97–111. https://doi.org/10.1002/jlb.40.1.97
Brown, R., Moore, L., Mani, A., Patel, S., Salinas, I. (2021) Effects of ploidy and salmonid alphavirus infection on the skin and gill microbiome of Atlantic salmon (Salmo salar). PLoS One. 16(2), e0243684.
Cámara-Ruiz, M., Cerezo, I., M., Guardiola, F., A. et al. (2021) Alteration of the immune response and the microbiota of the skin during a natural infection by Vibrio harveyi in European seabass (Dicentrarchus labrax). Microorganisms. 9(5), 964.
Carnevali, O., Maradonna, F. & Gioacchini, G. (2017) Integrated control of fish metabolism, wellbeing and reproduction: the role of probiotic. Aquaculture 472, 144-155. https://www.doi.org/10.1016/j.aquaculture.2016.03.037
Cerezuela, R., Meseguer, J. & Esteban, M. (2011) Current knowledge in synbiotic use for fish aquaculture: a review. J. Aquac. Res. Dev. 1, 1-7.
Cherrak, Y., Flaugnatti, N., Durand, E., Journet, L. & Cascales, E. (2019) Structure and Activity of the Type VI Secretion System. Microbiology spectrum, 7(4), 1.
Colorni, A., Avtalion, R., Knibb, W., Berger, E., J., Colorni, B. & Timan, B. (1998) Histopathology of sea bass (Dicentrarchus labrax) experimentally infected with Mycobacterium marinum and treated with streptomycin and garlic (Allium sativum) extract. Aquaculture. 160, 1-17.
Cordero, H., Guardiola, F., A., Tapia-Paniagua, S., T., Cuesta, A., Meseguer, J., Balebona, M., C., et al. (2015) Modulation of immunity and gut microbiota after dietary administration of alginate encapsulated Shewanella putrefaciens Pdp11 to gilthead seabream (Sparus aurata L.). Fish Shellfish Immunol. 45, 608-618. https://www.doi.org/10.1016/j.fsi.2015.05.010
Das, P., Mandal, S., Khan, A., Manna, S., K. & Ghosh, K. (2014) Distribution of extracellular enzyme-producing bacteria in the digestive tracts of 4 brackish water fish species. Turk. J. Zool. 38, 79-88. https://www.doi.org/10.3906/zoo-1205-3
Delcroix, J., Gatesoupe, F., J., Desbruyères, E., Huelvan, C., Le Delliou, H., Le Gall, M., M., Quazuguel, P., et al. (2015) The effects of dietary marine protein hydrolysates on the development of sea bass larvae, Dicentrarchus labrax, and associated microbiota. Aquac. Nutr. 21, 98-104. https://www.doi.org/10.1111/anu.12139
Duperron, S., Halary, S., Habiballah, M., Gallet, A., Huet, H., Duval, C. et al. (2019) Response of fish gut microbiota to toxin-containing cyanobacterial extracts: a microcosm study on the Medaka (Oryzias latipes). Environ SciTechnol Lett. 6, 341–7.
Egerton, S., Culloty, S., Whooley, J., Stanton, C. & Ross, R., P. (2018) The Gut Microbiota of Marine Fish. Frontiers in Microbiology. 9(873),. https://www.frontiersin.org/articles/https://doi.org/10.3389/fmicb.2018.00873/full
Elhassan, Y., Philp, A. & Lavery, G. (2017) Targeting NAD+ in Metabolic Disease: New Insights Into an Old Molecule. Journal of the Endocrine Society. 1(7), 816-835.
Estruch, G., Collado, M., Peñaranda, D., Vidal, A., T., Cerdá, M., J., Martínez, G., P., et al. (2015) Impact of fishmeal replacement in diets for gilthead sea bream (Sparus aurata) on the gastrointestinal microbiota determined by pyrosequencing the 16S rRNA gene. PLoS One 10, e0136389. https://www.doi.org/10.1371/journal.pone.0136389
Flerova, E. & Balabanova, L. (2013) Ultrastructure of granulocytes of teleost fish (Salmoniformes, Cypriniformes, Perciformes). Journal of Evolutionary Biochemistry and Physiology. 49(2), 223-233.
Fournier, V., Huelvan, C. & Desbruyeres, E. (2004) Incorporation of a mixture of plant feedstuffs as substitute for fish meal in diets of juvenile turbot (Psetta maxima). Aquaculture 236, 451-465. https://www.doi.org/10.1016/j.aquaculture.2004. 01.035
Gallet, A., Halary, S., Duval, C., Huet, H., Duperron, S. & Marie, B. (2023) Disruption of fish gut microbiota composition and holobiont’s metabolome during a simulated Microcystis aeruginosa (Cyanobacteria) bloom. Microbiome. 11, 108. https://doi.org/10.1186/s40168-023-01558-2
Ghanbari, M., Kneifel, W. & Domig, K., J. (2015) A new view of the fish gut microbiome: advances from next-generation sequencing. Aquaculture 448, 464-475. https://www.doi.org/10.1016/j.aquaculture.2015.06.033
Giatsis, C., Sipkema, D., Smidt, H. et al. (2015) The impact of rearing environment on the development of gut microbiota in tilapia larvae. Sci Rep. 5(1), 1-15.
Green, T., J., Smullen, R. & Barnes, A., C. (2013) Dietary soybean protein concentrate-induced intestinal disorder in marine farmed Atlantic salmon, Salmo salar is associated with alterations in gut microbiota. Vet. Microbiol. 166, 286-292. https://www.doi.org/10.1016/j.vetmic.2013.05.009
Hansen, G. & Olafsen, J. (1999) Bacterial interactions in early life stages of marine cold water fish. Microb. Ecol. 38, 1-26. https://www.doi.org/10.1007/s002489900158
Hlongwane, P., Mungra, N., Madheswaran, S., Akinrinmade, O., A., Chetty, S. & Barth, S. (2018) Human Granzyme B Based Targeted Cytolytic Fusion Proteins. Biomedicines. 6(2), 72.
Ho, J. & Kim, I. (2001) New species of Hatschekia Poche, 1902 (Copepoda: Hatschekiidae) parasitic on marine fishes of Kuwait. Syst Parasitol. 49, 73-79.
Hooper, L., V. & Gordon, J., I. (2001) Glycans as legislators of host-microbial interactions: spanning the spectrum from symbiosis to pathogenicity. Glycobiology 11, 1R-10R. https://doi.org/10.1093/glycob/11.2.1R
Hovanec, T. (2019) Dr. How to harness bacteria to cycle your saltwater tank quickly! | MACNA 2019. BrsTV. https://www.youtube.com/watch?v=zDI7sxqC-ss
James, A., G. (1988) Are clupeid microphagists herbivorous or omnivorous? A review of the diets of some commercially important clupeids. S. Afr. J. Mar. Sci. 7, 161-177. https://www.doi.org/10.2989/025776188784379017
Kapoor, B. & Khawna, B. (1993) The potential spectrum of the gut in teleost fishes. Adv. Fish Res. 1, 221-226.
Karachle, P., K. & Stergiou, K., I. (2010) Gut length for several marine fish: relationships with body length and trophic implications. Mar. Biodivers. Rec. 3, e106. https:///www.doi.org/10.1017/S1755267210000904
Kelly, C. & Salinas, I. (2017) Under pressure: Interactions between commensal microbiota and the teleost immune system. Front. Immunol. 8, 1.
Kim, B.,-R., Shin. J., Guevarra, R., B., et al. (2017) Deciphering diversity indices for a better understanding of microbial communities. J Microbiol Biotechnol. 27(12), 2089-2093.
Kim, J. & Lee, J., L. (2017) Correlation of Total Bacterial and Vibrio spp. Populations between Fish and Water in the Aquaculture System. Frontiers in Marine Science. 4,.
Kiron, V. (2012) Fish immune system and its nutritional modulation for preventive health care. Anim. Feed Sci. Technol. 173, 111-133. https://www.doi.org/10.1016/j.anifeedsci.2011.12.015
Kline, D., I., Kuntz, N., M., Breitbart, M., Knowlton, N. & Rohwer, F. (2006) Role of elevated organic carbon levels and microbial activity in coral mortality. Marine Ecology Progress Series. 314, 119-125.
Langille, M., G., Zaneveld, J., Caporaso, J., G. et al. (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol. 31(9), 814-821.
Larsen, A., M. (2014) Studies on the Microbiota of Fishes and the Factors Influencing Their Composition. Auburn, AL: Auburn University.
Lasica, A., Ksiazek, M., Madej, M. & Potempa, J. (2017) The Type IX Secretion System (T9SS): Highlights and Recent Insights into Its Structure and Function. Frontiers in Cellular and Infection Microbiology. 7,.
Legrand, T., P., R., A., Catalano, S., R., Wos-Oxley, M., L., Stephens, F., Landos, M., Bansemer, M., S., Stone, D., A., J., Qin, J., G. & Oxley, A., P., A. (2018) The Inner Workings of the Outer Surface: Skin and Gill Microbiota as Indicators of Changing Gut Health in Yellowtail Kingfish. Front. Microbiol. 8:2664. https://www.doi.org/10.3389/fmicb.2017.02664
Lesel, R., De La Noüe, J. & Choubert, G. (1989) Fecal bacterial flora of rainbow trout under antibiotic treatment: effect of the number of pyloric caeca and the lipid content of food. Aquaculture: A Biotechnology in Progress, Vol. 1. De Pauw, N., Jaspers, E., Ackefors, H. & Wilkins, N. (eds.). Bredene: European Aquaculture Society, 592.
Li, P. & Gatlin, D., M. (2003) Evaluation of brewers yeast (Saccharomyces cerevisiae) as a feed supplement for hybrid striped bass (Morone chrysopsX—M. saxatilis). Aquaculture. 219(1), 681-692.
Li, T., Li, H., Gatesoupe, F-J. et al. (2017) Bacterial signatures of ‘red-operculum’ disease in the gut of crucian carp (Carassius auratus). Microb Ecol. 74(3), 510-521.
Liu, Q., Lai, Z., Gao, Y. et al. (2021) Connection between the gut microbiota of largemouth bass (Micropterus salmoides) and microbiota of the pond culture environment. Microorganisms. 9(8), 1770.
Llewellyn, M., Leadbeater, S., Garcia, C. et al. (2017) Parasitism perturbs the mucosal microbiome of Atlantic Salmon. Sci Rep. 7(1), 1-10.
Llewellyn, M., S., Boutin, S., Hoseinifar, S., H. & Derome, N. (2018) Teleost microbiomes: the state of the art in their characterization, manipulation and importance in aquaculture and fisheries. Front Microbiol. 2014, 5. http://journal.frontiersin.org/article/10.3389/fmicb.2014.00207/abstract
Lobo, C., Moreno-Ventas, X., Tapia-Paniagua, S., Rodríguez, C., Moriñigo, M., A. & de La Banda, I., G. (2014) Dietary probiotic supplementation (Shewanella putrefaciens Pdp11) modulates gut microbiota and promotes growth and condition in Senegalese sole larviculture. Fish Physiol. Biochem. 40, 295-309. https://www.doi.org/10.1007/s10695-013-9844-0
Lom, J. & Corliss, J. (1971) Morphogenesis and Cortical Ultrastructure of Brooklynella hostilis, a Dysteriid Ciliate Ectoparasitic on Marine Fishes. The Journal of Eukaryotic Microbiology. 18(2), 261-281.
López Nadal, A., Peggs, D., Wiegertjes, G., F. & Brugman, S. (2018) Exposure to Antibiotics Affects Saponin ImmersionInduced Immune Stimulation and Shift in Microbial Composition in Zebrash Larvae. Front Microbiol 9, 2588. https://doi.org/10.3389/fmicb.2018.02588
López-Dóriga, M., Barnes, A., dos Santos, N. & Ellis, A. (2000) Invasion of fish epithelial cells by Photobacterium damselae subsp. piscicida: evidence for receptor specificity, and effect of capsule and serum. Microbiology. 146(1), 21-30.
Lorgen-Ritchie, M., Clarkson, M., Chalmers, L., Taylor, J., F., Migaud, H. & Martin, S., A. M. (2021) A temporally dynamic gut microbiome in Atlantic salmon during freshwater recirculating aquaculture system (RAS) production and post-seawater transfer. Front. Mar. Sci. 8, 869. https://doi.org/10.3389/fmars.2021.711797
Lorgen-Ritchie, M., Webster, T., U., McMurtrie, J., Bass, D., Tyler, C., R., Rowley, A. & Martin, S., A., M. (2023) Microbiomes in the context of developing sustainable intensified aquaculture. Frontiers in Microbiology. 14, 1200997
Ma, C., Chen, C., Jia, L., He, X. & Zhang, B. (2019) Comparison of the intestinal microbiota composition and function in healthy and diseased Yunlong Grouper. AMB Express. 9(1), 1-11.
Martins, J. & Vasconcelos, V. (2015) Cyanobactins from cyanobacteria: current genetic and chemical state of knowledge. Mar Drugs. 13, 6910-6946.
Mashoof, S. & Criscitiello, M., F. (2016) Fish Immunoglobulins. Biology. 5(4), 45.
McBride, M., J. & Nakane, D. (2015) Flavobacterium gliding motility and the type IX secretion system. Curr. Opin. Microbiol. 28, 72-77.
Merrifield, D., L. & Rodiles, A. (2015) The fish microbiome and its interactions with mucosal tissues. Mucosal Health in Aquaculture. Peatman, E. (ed.) Academic Press, San Diego, CA. pp 273-295.
Mihalitsis, M. & Bellwood, D. (2017) A morphological and functional basis for maximum prey size in piscivorous fishes. PLoS ONE. 12(9),.
Miyake, S., Ngugi, D. K. & Stingl, U. (2015) Diet strongly influences the gut microbiota of surgeonfishes. Mol. Ecol. 24, 656-672. https://www.doi.org/10.1111/mec.13050
Miyake, S., Soh, M., Azman, M., N., Ngoh, S., Y., Orbán, L., Seedorf, H. (2020) Insights into the microbiome of farmed Asian sea bass (Lates cal-carifer) with symptoms of tenacibaculosis and description of Tenacibaculum singaporense sp. nov. Antonie Van Leeuwenhoek. 113(6), 737-752.
Monroig, Ó., Tocher, D., R. & Navarro, J., C. (2013) Biosynthesis of polyunsaturated fatty acids in marine invertebrates: recent advances in molecular mechanisms. Mar. Drugs 11, 3998-4018. https://www.doi.org/10.3390/md11103998
Moon, D. (2021) Boosting NAD+ to Reverse Aging? Overview of NR and NMN. GeneticLifeHacks.com. https://www.geneticlifehacks.com/nad-reversing-aging-overview-of-nr-and-nmn/
Mouchet, M., A., Bouvier, C., Bouvier, T., Troussellier, M., Escalas, A. & Mouillot, D. (2012) Genetic difference but functional similarity among fish gut bacterial communities through molecular and biochemical fingerprints. FEMS Microbiol. Ecol. 79, 568-580. https://www.doi.org/10.1111/j.1574-6941.2011. 01241.x
Mougin, J. & Joyce, A. (2022) Reviews in Aquaculture. Fish disease prevention via microbial dysbiosis-associated biomarkers in aquaculture. 15, 579-594. https://doi.org/10.1111/raq.12745
Muñoz-Baquero, M., Lorenzo-Rebenaque, L., Garc´ıa-Va´zquez, F., A., Garc´ıa-Pa´rraga, D., Mart´ınez-Priego, L., De Marco-Romero, G., Gala´n-Vendrell, I., D’Auria, G. & Marco-Jime´nez, F. (2023) Unveiling Microbiome Signature In Inner Body Fluids: Comparison Between Wild And Aquarium Small-Spotted Catshark (Scyliorhinus canicular). Frontiers in Marine Science. https://www.doi.org/10.3389/fmars.2023.1151119
Murdoch, C., C. & Rawls, J., F. (2019) Commensal microbiota regulate vertebrate innate immunity-insights from the zebrafish. Front. Immunol. 10:2100. https://doi.org/10.3389/fimmu.2019.02100
Navarrete, P., Espejo, R. T. & Romero, J. (2009) Molecular analysis of microbiota along the digestive tract of juvenile Atlantic salmon (Salmo salar L.). Microb. Ecol. 57, 550-561. https://www.doi.org/10.1007/s00248-008-9448-x
Naya-Català, F., Piazzon, M., C., Calduch-Giner, J. A., Sitjà-Bobadilla, A. & Pérez-Sánchez, J. (2022) Diet and host genetics drive the bacterial and fungal intestinal metatranscriptome of Gilthead Sea bream. Front. Microbiol. 13, 883738. https://doi.org/10.3389/fmicb.2022.883738
Nayak, S., K. (2010) Role of gastrointestinal microbiota in fish. Aquac. Res. 41, 1553-1573. https://www.doi.org/10.1111/j.1365-2109.2010.02546.x
Neuman, C., Hatje, E., Zarkasi, K. Z., Smullen, R., Bowman, J., P. & Katouli, M. (2016) The effect of diet and environmental temperature on the faecal microbiota of farmed Tasmanian Atlantic Salmon (Salmo salar L.). Aquac. Res. 47, 660-672. https://www.doi.org/10.1111/are.12522
Noga, E., J. (2010) Fish Disease. Diagnosis and Treatment. Second Edition. Wiley-Blackwell, John Wiley & Sons Inc. p 139.
Oliva-Teles, A. & Goncalves, P. (2001) Partial replacement of fishmeal by brewers yeast (Saccaromyces cerevisae) in diets for sea bass (Dicentrarchus labrax) juveniles. Aquaculture. 202(3), 269-278.
Orlandi, I., Alberghina, L. & Vai, M. (2020) Nicotinamide, Nicotinamide Riboside and Nicotinic Acid-Emerging Roles in Replicative and Chronological Aging in Yeast. Biomolecules. 10(4), 604.
Ortiz-Estrada, A., M., Gollas-Galván, T., Martínez-Cordova, L., R. & Martínez-Porchas, M. (2019) Predictive functional profiles using metagenomic 16S rRNA data: a novel approach to understanding the microbial ecology of aquaculture systems. Rev Aquacult. 11(1), 234-245.
Perdiguero, P., Martin-Martin, A., Benedicenti, O., Diaz-Rosales, P., Morel, E., Munoz-Atienza, E., Garcia-Flores, M., Simon, R., Soleto, I., Cerutti, A. & Tafalla, C. (2019) Teleost IgD+Ig- B Cells Mount Clonally Expanded and Mildly Mutated Intestinal IgD Responses in the Absence of Lymphoid Follicles. Cell Reports. 29(13), 4223-4235.
Pérez-Pascual, D., Lunazzi, A., Magdelenat, G., Rouy, Z., Roulet, A., Lopez-Roques, C., Larocque, R., Barbeyron, T., Gobet, A., Michel, G., Bernardet, J. & Duchaud, E. (2017) The Complete Genome Sequence of the Fish Pathogen Tenacibaculum maritimum Provides Insights into Virulence Mechanisms. Frontiers in Microbiology. 8,.
Reid, K., M., Patel, S., Robinson, A., J., et al. (2017) Salmonid alphavirus infection causes skin dysbiosis in Atlantic salmon (Salmo salar L.) post-smolts. PLoS One. 12(3):e0172856.
Richardson, L., L., Sekar, R., Myers, J., L., Gantar, M., Voss, J., D., Kaczmarsky, L., Remily, E., R., Boyer, G., L. & Zimba, P., V. (2007) The presence of the cyanobacterial toxin microcystin in black band disease of corals. FEMS Microbiology Letters. 272(2), 182-187.
Riley, M., A. & Wertz, J., E. (2002) Bacteriocin diversity: ecological and evolutionary perspectives. Biochimie. 84, 357-364.
Ringø, E., Lødemel, J., B., Myklebust, R., Jensen, L., Lund, V., Mayhew, T., M., et al. (2002) The effects of soybean, linseed and marine oils on aerobic gut microbiota of Arctic charr Salvelinus alpinus L. before and after challenge with Aeromonas salmonicida ssp. salmonicida. Aquac. Res. 33, 591-606. https://www.doi.org/10.1046/j.1365-2109.2002.00697.x
Ringø, E., Zhou, Z., Vecino, J., L., G., Wadsworth, S., Romero, J., Krogdahl, Å., et al. (2016) Effect of dietary components on the gut microbiota of aquatic animals. A never-ending story? Aquac. Nutr. 22, 219-282. https://www.doi.oeg/10.1111/anu.12346
Ritchie, K., B. & Smith, G., W. (1995) Preferential carbon utilization by surface bacterial communities from water mass, normal, and white-band diseased Acropora cervicornis. Mol Mar Biol Biotechnol. 4, 345-354.
Rohde, K. (1932) Ciliophora (ciliates). Marine Parasitology. Rohde, K. (ed.). CSIRO Publishing, Clayton, Australia.
Rombout, J., H., Abelli, L., Picchietti, S., Scapigliati, G. & Kiron, V. (2011) Teleost intestinal immunology. Fish Shellfish Immunol. 31, 616-626. https://www.doi.org/10.1016/j.fsi. 2010.09.001
Rosado, D., Pérez-Losada, M., Pereira, A., Severino, R. & Xavier, R. (2021) Effects of aging on the skin and gill microbiota of farmed seabass and seabream. Anim Microbiome. 3(1), 1-14.
Rosado, D., Pérez-Losada, M., Severino, R. & Xavier, R. (2022) Monitoring infection and antibiotic treatment in the skin microbiota of farmed European seabass (Dicentrarchus Labrax) fingerlings. Microb Ecol. 83(3), 789-797.
Rosado, D., Perez-Losada, M., Severino, R., Cable, J. & Xavier, R. (2019b) Characterization of the skin and gill microbiomes of the farmed seabass (Dicentrarchus labrax) and seabream (Sparus aurata). Aquaculture. 500, 57-64.
Rosado, D., Xavier, R., Severino, R., Tavares, F., Cable, J. & Pérez-Losada, M. (2019a) Effects of disease, antibiotic treatment and recovery trajectory on the microbiome of farmed seabass (Dicentrarchus labrax). Sci Rep. 9(1), 1-11.
Rozas-Serri, M. (2019) Gill diseases in marine salmon aquaculture with an emphasis on amoebic gill disease. CAB Reviews Perspectives in Agriculture Veterinary Science Nutrition and Natural Resources. 14, 1-15.
Santoro, E., P., Borges, R., M., Espinoza, J., L., Freire, M., Messias, C., S., M., A., Villela, H., D., M., Pereira, L., M., Vilela, C., L., S., Rosado, J., G., Cardoso, P., M., Rosado, P., M., Assis, J., M., Duarte, G., A., S., Perna, G., Rosado, A., S., Macrae, A., Dupont, C., L., Nelson, K., E., Sweet, M., J., Voolstra, C., R. & Peixoto, R., S. (2021) Coral microbiome manipulation elicits metabolic and genetic restructuring to mitigate heat stress and evade mortality. Science advances. 7(33), 3088.
Santos, F., F., Yamamoto, D., Abe, C., M., Bryant, J., A., Hernandes, R., T., Kitamura, F., C., Castro, F., S., Valiatti, T., B., Piazza, R., Elias, W., P., Henderson, I., R. & Gomes, T. (2019) The Type III Secretion System (T3SS)-Translocon of Atypical Enteropathogenic Escherichia coli (aEPEC) Can Mediate Adherence. Frontiers in microbiology. 10, 1527.
Scapigliati, G., Fausto, A., M. & Picchietti, S. (2018) Fish Lymphocytes: An Evolutionary Equivalent of Mammalian Innate-Like Lymphocytes? Frontiers in immunology. 9, 971.
Segata, N., Izard, J., Waldron, L., et al. (2011) Metagenomic biomarker discovery and explanation. Genome Biol. 12(6), 1-18.
Shabir, U., Ali, S., Magray, A., Ganai, B., Firdous, P., Hassan, T. & Nazir, R. (2018) Fish antimicrobial peptides (AMP’s) as essential and promising molecular therapeutic agents: A review. Microbial pathogenesis. 114, 50-56.
Sila, A., Nedjar-Arroume, N., Hedhili, K., Chataigné, G., Balti, R., Nasri, M., et al. (2014) Antibacterial peptides from barbel muscle protein hydrolysates: activity against some pathogenic bacteria. LWT Food Sci. Technol. 55, 183-188. https://www.doi.org/10.1016/j.lwt.2013.07.021
Siriyappagouder, P., Galindo-Villegas, J., Dhanasiri, A., K., S., Zhang, Q., Mulero, V., Kiron, V. et al. (2020) Pseudozyma priming influences expression of genes involved in metabolic pathways and immunity in zebrafish larvae. Front. Immunol. 11, 987. https://doi.org/10.3389/fimmu.200.00978
Siriyappagouder, P., Kiron, V., Lokesh, J., Rajeish, M., Kopp, M. & Fernandes, J. (2018) The intestinal mycobiota in wild zebrafish comprises mainly Dothideomycetes while Saccharomycetes predominate in their laboratory-reared counterparts. Front. Microbiol. 9, 387. https://doi.org/10.3389/fmicb.2018.00387
Sitjà-Bobadilla, A., Gil-Solsona, R., Estensoro, I., Piazzon, M., Martos-Sitcha, J., Picard-Sánchez, A., Fuentes, J., Sancho, J., Calduch-Giner, J., Hernández, F. & Pérez-Sánchez, J. (2019) Disruption of gut integrity and permeability contributes to enteritis in a fish-parasite model: a story told from serum metabolomics. Parasites & Vectors. 12,.
Skrodenyte-ArbaCIauskiene, V. (2007) Enzymatic activity of intestinal bacteria in roach Rutilus rutilus L. Fish. Sci. 73, 964-966. https://www.doi.org/10.1111/j.1444-2906.2007. 01421.x
Slinger, J., Adams, M., B., Stratford, C., N., Rigby, M. & Wynne, J., W. (2021) The effect of antimicrobial treatment upon the gill bacteriome of Atlantic salmon (Salmo salar L.) and progression of amoebic gill disease (AGD) in vivo. Microorganisms. 9, 987. https://doi.org/10.3390/microorganisms9050987
Spanova, M. & Daum, G. (2011) Squalene–biochemistry, molecular biology, process biotechnology, and applications. Eur J Lipid Sci Technol. 113(11), 1299-1320.
Spor, A., Koren, O. & Ley, R. (2011) Unravelling the effects of the environment and host genotype on the gut microbiome. Nat. Rev. Microbiol. 9, 279-290. https://doi.org/10.1038/nrmicro2540
Stevens, C., E. & Hume, I., D. (2004) Comparative Physiology of the Vertebrate Digestive System. Cambridge: Cambridge University Press.
Stevens, J., Jackson, R., Olson, J. (2016) Bacteria associated with lionsh (Pterois volitans/miles complex) exhibit antibacterial activity against known fish pathogens. Mar Ecol Prog Ser 558, 167–180. https://doi.org/10.3354/meps11789
Sullam, K., E., Essinger, S., D., Lozupone, C., A., O’Connor, M., P., Rosen, G., L., Knight, R., et al. (2012) Environmental and ecological factors that shape the gut bacterial communities of fish: a meta-analysis. Mol. Ecol. 21, 3363-3378. https://www.doi.org/10.1111/j.1365-294X.2012.05552.x
Swiatecka, D., Markiewicz, L., H. & Wróblewska, B. (2012) Pea protein hydrolysate as a factor modulating the adhesion of bacteria to enterocytes, epithelial proliferation and cytokine secretion–an in vitro study. Cent. Eur. J. Immunol. 37, 227-231. https://www.doi.org/10.5114/ceji.2012.3079
Tapia-Paniagua, S., T., Ceballos-Francisco, D., Balebona, M., C., Esteban, M., A. & Moriñigo, M., A. (2018) Mucus glycosylation, immunity and bacterial microbiota associated to the skin of experimentally ulcered gilthead seabream (Sparus aurata). Fish Shellfish Immunol. 75, 381-390.
Thiéry, R., Cozien, J., Cabon, J., Lamour, F., Baud, M. & Schneemann, A. (2006) Induction of a Protective Immune Response against Viral Nervous Necrosis in the European Sea Bass Dicentrarchus labrax by Using Betanodavirus Virus-Like Particles. Journal of Virology. 80(20), 10201.
Torrecillas, S., Montero, D. & Izquierdo, M. (2014) Improved health and growth of fish fed mannan oligosaccharides: potential mode of action. Fish Shellfish Immunol. 36, 525-544. https://doi.org/10.1016/j.fsi.2013.12.029
Tretina, K., Park, E., S., Maminska, A. & MacMicking, J., D. (2019) Interferon-induced guanylate-binding proteins: Guardians of host defense in health and disease. The Journal of Experimental Medicine. 216(3), 482-500.
Valero, Y., Saraiva‐Fraga, M., Costas, B. & Guardiola, F., A. (2019) Antimicrobial peptides from fish: beyond the fight against pathogens. Reviews in Aquaculture. 12, 224-253.
Vayssier-Taussat, M., Albina, E., Citti, C., et al. (2014) Shifting the paradigm from pathogens to pathobiome: new concepts in the light of metaomics. Front Cell Infect Microbiol. 4, 29.
Vestrum, R., I., Forberg, T., Luef, B., Bakke, I., Winge, P., Olsen, Y., et al. (2022) Commensal and opportunistic bacteria present in the microbiota in Atlantic cod (Gadus morhua) larvae differentially alter the hosts’ innate immune responses. Microorganisms. 10, 24. https://doi.org/10.3390/microorganisms10010024
Walke, J., B., et al. (2017) Dominance-function relationships in the amphibian skin microbiome. Environ. Microbiol. 19, 3387–3397.
Wang, S. & Loreau, M. (2014) Ecosystem stability in space: α, β and γ variability. Ecol Lett. 17(8), 891-901.
Webster, T., M., U., Rodriguez-Barreto, D., Consuegra, S. & Garcia de Leaniz, C. (2019) Cortisol-induced signatures of stress in the fish microbiome. bioRxiv:826503. https://doi.org/10.1101/826503
Wilson, J., M. & Castro, L., F., C. (2010) Morphological diversity of the gastrointestinal tract in fishes. Fish Physiol. 30, 1-55. https://www.doi.org/10.1016/S1546-5098(10)03001-3
Wilson, M. World Feeds Limited, 3b Coulman Street Industrial Estate, Thorne, Doncaster, DN8 5JS, United Kingdom.
Wu, J., Mao, C., Deng, Y., et al. (2019) Diversity and abundance of antibiotic resistance of bacteria during the seedling period in marine sh cage-culture areas of Hainan, China. Mar Pollut Bull 141:343–349. https://doi.org/10.1016/j.marpolbul.2019.02.069
Xavier, R., Pereira, A., Pagan, A., Hendrick, G., C., Nicholson, M., D., Rosado, D., Soares, M., C., Pérez-Losada, M., Sikkel, P., C. (2020) The effects of environment and ontogeny on the skin microbiome of two Stegastes damselfishes (Pomacentridae) from the eastern Caribbean Sea. Mar Biol. 167(7),1-12.
Xu, Z., Takizawa, F., Casadei, E., Shibasaki, Y., Ding, Y., Sauters, T., J., C., et al. (2020) Specialization of mucosal immunoglobulins in pathogen control and microbiota homeostasis occurred early in vertebrate evolution. Sci. Immunol. 5, 3254. https://doi.org/10.1126/sciimmunol.aay3254
Yatsunenko, T., Rey, F., E., Manary, M., J., Trehan, I., Dominguez-Bello, M., G., Contreras, M., et al. (2012) Human gut microbiome viewed across age and geography. Nature 486, 222-227. https://www.doi.org/10.1038/nature11053
Yoon, J., Matsuo, Y., Matsuda, S., Adachi, K., Kasai, H. & Yokota, A. (2018) Rubritalea sabuli sp. nov., a carotenoid-and squalene-producing member of the family Verrucomicrobiaceae, isolated from marine sediment. Int J Syst Evol Microbiol. 58(4), 992-997.
Yoshimizu, M., Kimura, T. & Sakai, M. (1980) Microflora of the embryo and the fry of salmonids. Bull. Jpn. Soc. Sci. Fish. 46, 967-975. https://www.doi.org/10.2331/suisan.46.967
Yu, Y., Wang, Q., Huang, Z., Ding, L. & Xu, Z. (2020) Immunoglobulins, Mucosal Immunity and Vaccination in Teleost Fish. Frontiers in immunology. 11, 567941.
Zanefeld, J., R. et al. (2017) Stress and stability: applying the Anna Karenina principle to animal microbiomes. Nat. Microbiol. 2, 17121.
Zarkasi, K., Z., Taylor, R., S., Abell, G., C., Tamplin, M., L., Glencross, B., D. & Bowman, J., P. (2016) Atlantic salmon (Salmo salar L.) gastrointestinal microbial community dynamics in relation to digesta properties and diet. Microb. Ecol. 71, 589-603. https://www.doi.org/10.1007/s00248-015-0728-y
Zeng, A., Tan, K., Gong, P. et al. (2020) Correlation of microbiota in the gut of fish species and water. 3 Biotech. 0(11), 1-10.
Zhang, L., Ni, C., Xu, W., Dai, T., Yang, D., Wang, Q., Zhang, Y. & Liu, Q. (2016) Intramacrophage Infection Reinforces the Virulence of Edwardsiella tarda. Journal of bacteriology. 198(10), 1534-1542.
Zhang, M., Shan, C., Tan, F., Limbu, S., M., Chen, L. & Du, Z. (2020) Gnotobiotic models: powerful tools for deeply understanding intestinal microbiota-host interactions in aquaculture. Aquaculture 517:734800. https://doi.org/10.1016/j.aquaculture.2019.734800
Zhang, X., Ding, L., Yu, Y. et al. (2018) The change of teleost skin commensal microbiota is associated with skin mucosal transcriptomic responses during parasitic infection by Ichthyophthirius multifillis. Front Immunol. 9, 2972.
Zhang, Y. & Gui, J. (2012) Molecular regulation of interferon antiviral response in fish. Developmental and comparative immunology. 38(2), 193-202.
Zhou, Z., Liu, Y., Shi, P., He, S., Yao, B. & Ringø, E. (2009) Molecular characterization of the autochthonous microbiota in the gastrointestinal tract of adult yellow grouper (Epinephelus awoara) cultured in cages. Aquaculture 286, 184-189. https://www.doi.org/10.1016/j.aquaculture.2008.10.002
Zhou, Z., Yao, B., Romero, J., Waines, P., Ringø, E., Emery, M., et al. (2014) Methodological approaches used to assess fish gastrointestinal communities. Aquaculture Nutrition: Gut Health, Probiotics and Prebiotics, Merrifield, D. & Ringø, E. (eds.). John Wiley & Sons Ltd., Hoboken, NJ.



0 Comments