
Maintaining coral coloration is the goal of many reef hobbyists, but is often a source of frustration. Most hobby literature I’ve seen concerning the maintenance of coral coloration in captivity involves lighting (an important consideration!) but there is more to the story. This article will examine (according to latest theories) why light promotes coloration and how to increase your chances of success in maintaining these colors.
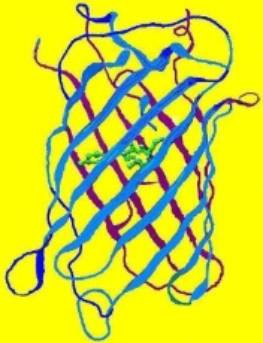
Figure 1. The structure of a fluorescent protein -it is the basis of our understanding of how various factors can affect coral coloration. Drawing by the author.
This time, we’ll look at colors of some of the most popular reef aquarium corals – Acropora species (Family Acroporidae). There is a considerable amount of relatively new information presented in this article.
I have found that, in general, there are two types of hobbyists. One group – the majority, in my opinion, wants an easy answer to how to maintain color, and doesn’t particularly care about the ‘why’. On the other hand, a few have an insatiable appetite for every morsel of minutiae. Hence, I get comments about a particular article I’ve written ranging from ‘too technical’ to ‘not enough details’. If you’re in the former group, skip to the ‘In Closing’ section at the end of this article (although I would hope that you would at least scroll through the bullet points and should certainly be familiar with the Glossary contained within Step One, below). If you’re of the latter group, the details contained below will fall into place, and you’ll have a good understanding as to ‘why’ and ‘how’.
The Fluorescent Protein
Our understanding of coral coloration begins with the basic unit – the fluorescent protein. Originally isolated from the jellyfish (Aequoria victoria), the group of Green Fluorescent Proteins (GFPs) has expanded and now includes a number of taxa, many of them Anthozoans. However, the basic structure of the protein remains remarkably similar (see Figure 1, above). Bands (called staves) of proteins form a protective barrel around a fluorescent nucleus (called a fluorophore, shown as a green internal structure in the drawing). If the protein is not fluorescent (or very weakly so), this nucleus is called a chromophore. The individual amino acids forming these proteins usually number around 250. Some of these amino acids are charged, and these very weak ‘magnets’ tend to hold the structure intact. However, various factors (pH, light, ultraviolet radiation, infrared radiation, metal ions, etc.) can alter the relationship between the charge of amino acids and the fluorophore/chromophore, thus causing it to twist (called a ‘hula twist’). In some cases, this causes an irreversible color change (perhaps from green to red), make the chromophore become a fluorescent fluorophore, cause fluorescence to vanish completely (in at least one case), and so on. The concept that the protein structure can be altered by external stimuli is an important one.
Our journey through the jungle of details begins.
Glossary
The following terms will be used in this article:
- Absorbance
- Ability of a solution or layer of a substance to retain light without reflection or transmission.
- Absorption
- The process in which incident radiation is retained without reflection or transmission.
- Clade
- For our purposes in this article, a grouping of pigments based on similar features inherited from a common ancestor. Pigments from corals includes Clades A, B, C (including sub-clades C1, C2 and C3), and D. Clades can refer to living organisms as well (clades of Symbiodinium – zooxanthellae – are a good example.)
- Chromophore
- The colorful but non-fluorescent portion of a pigment molecule. In some cases, chromophore refers to a granular packet containing many pigment molecules.
- Emission
- That light which is fluoresced by a fluorescent pigment.
- Excitation
- That light absorbed by a fluorescent pigment. Some of the excitation light is fluoresced or emitted at a less energetic wavelength (color).
- Fluorescence
- Absorption of radiation at one wavelength (or color) and emission at another wavelength (color). Absorption is also called excitation. Fluorescence ends very soon after the excitation source is removed (on the order of ~2-3 nanoseconds: Salih and Cox, 2006).
- Fluorophore
- A fluorescent portion of a pigment molecule.
- ‘Hula Twist’
- A bending of a pigment molecule resulting in a change of apparent color. Molecular bonds are not broken; therefore the pigment can shift back and forth, with movements reminiscent of a hula dancer.
- Photobleaching
- Some pigments, such as Dronpa, loss fluorescence if exposed to strong light (in this case, initially appearing green and bleaching to a non-fluorescent state when exposed to blue-green light). Photobleaching can obviously cause drastic changes in apparent fluorescence. In cases where multiple pigments are involved, the loss of fluorescence (or energy transfer from a donor pigment to an acceptor pigment) could also result in dramatic shifts in apparent color.
- Photoconversion
- A rearrangement of the chemical structure of a colorful protein by light. Depending upon the protein, photoconversion can increase or decrease fluorescence (in processes called photoactivation and photobleaching, respectively). Photoconversion can break proteins’ molecular bonds (as with Kaede and Eos fluorescent pigments) resulting in an irreversible color shift, or the molecule can be ‘twisted’ by light energy (a ‘hula twist’) where coloration reversal are possible depending upon the quality or quantity of light available. This process is known as photoswitching).
- Threshold or Coloration Threshold
- The point at which pigment production is sufficient to make its fluorescence (or in the case of non-fluorescent chromoproteins, it absorption) visually apparent. The term threshold generally refers pigment production, although, in some cases, it could apply to a light level where a pigment disappears (as in the cases of photobleaching, or photoconversion).
- Up-regulation
- Genes containing information such as coloration can sometimes be turned on (up-regulated) or off (down-regulated) by external stimuli such as light.
Step One: Identify fluorescent proteins
To start, we will examine different ways of identifying fluorescent proteins. The most obvious way is to classify them by color. In the genus Acropora, there are four basics types of colorful proteins. These are:
Cyan, Green, and Red Fluorescent Proteins and Non-fluorescent (or very weakly fluorescent) Chromoproteins. For our purposes, the coral glows when illuminated with an actinic lamp or blue LEDs, the protein is fluorescent. If not, it is a chromoprotein.
To further define these proteins, we’ll use these definitions:
Cyan Fluorescent Protein (CFP): Blue-green pigments with fluorescent emissions in the range of ~477-500nm (a definition used by researchers, although proteins with emissions into the violet portion of the spectrum have been noted). Cyan and green pigments share a similar chromophore structure. See Figures 2 and 3.
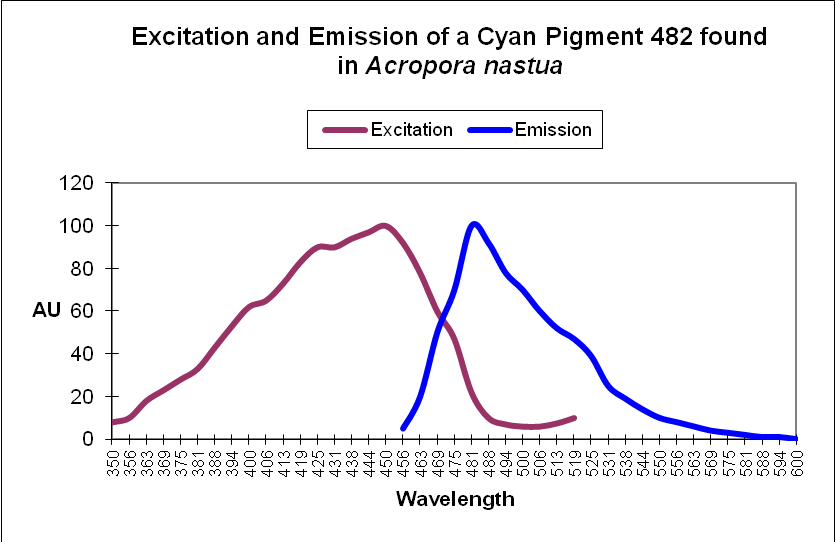
Figure 2. This cyan fluorescing protein absorbs blue light and fluoresces in the blue-green portion of the spectrum. The shape of the emission is fairly typical for a cyan fluorescent protein.
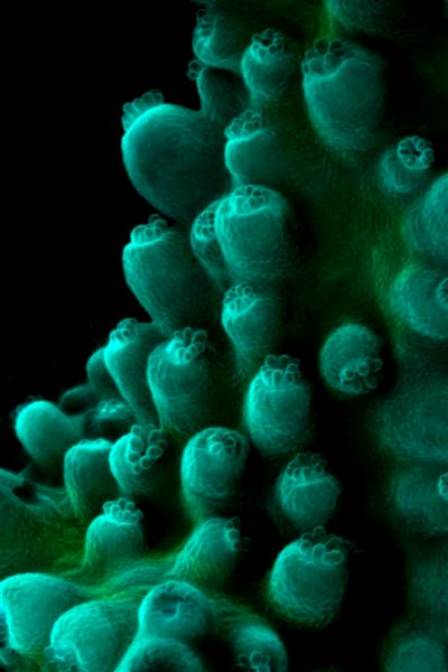
Figure 3. Cyan fluorescence of an unidentified Acropora. A bit of green fluorescent protein is also present.
Green Fluorescent Protein (GFP): Fluorescent pigments with emissions of 500-525nm. See Figures 4 and 5.
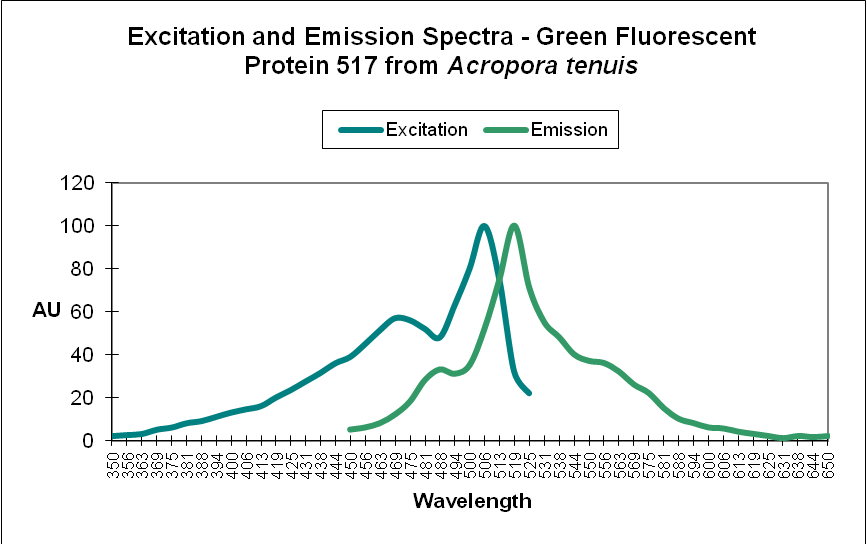
Figure 4. A Green Fluorescent Protein (GFP) from Acropora tenuis. Note the double peaks in both excitation and emission wavelengths. Does this protein change color from blue-green to green as it matures? See Figure 21 for another example from an Acropora specimen. Not all GFP emissions have these double peaks. After Papina et al., 2002.
DsRed-type pigment: A type of red fluorescent pigment with a single primary emission bandwidth at 574-630nm. Originally found in the false coral Discosoma, many other corals (including Acropora species) may contain orange/red DsRed-type proteins.See Figures 6 and 7.
Figure 7. Red fluorescence of a DsRed-type protein in an Acropora (A. millepora?). Photo by Justin Miedwig.
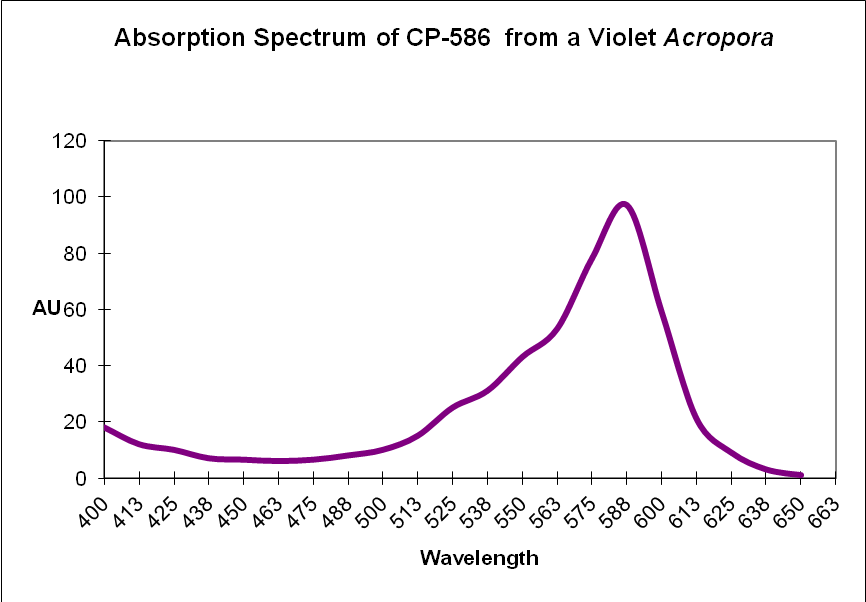
Chromoprotein pigment (CP): A non-fluorescent but colorful pigment. These pigments appear colorful because they preferentially absorb some portions of the spectrum and reflect others. For example, a chromoprotein with a maximum absorption at 580nm might appear purple because it reflects blue and red wavelengths. Generally, chromoproteins are purple, blue, mauve, or pink. ‘Pocilloporan’ is a generic term for non-fluorescent coral proteins. See Figures 8 and 9.
Figure 8. The purple-blue coloration in this Acropora is due to the presence of a non-fluorescent chromoprotein. Zooxanthellae within the polyps apparently lack colorful proteins and protection from excessive light. Photo by the author.
Figure 9. Recall that chromoproteins absorb some wavelengths and reflect others. In this case, a violet Acropora absorbs some green, yellow, orange and a few red wavelengths. It strongly reflects violet/blue and far-red light, making it appear violet.
Step 2: Compile a List of Known Acropora Proteins (listed by absorption for chromoproteins, and emission for fluorescent proteins).
See Reference list of about 160 at the end of this article. CP = non-fluorescent protein. CFP = Cyan fluorescent protein. GFP = Green Fluorescent Protein. DsRed = Discosoma Red- type fluorescent protein.
| Emission/Abs | Host | Type |
|---|---|---|
| CP-580 | Acropora aculeus | Chromoprotein |
| CP-580 | Acropora aculeus | Chromoprotein |
| CP-500 | Acropora sp. | Chromoprotein |
| CP-578 | Acropora digitifera | Chromoprotein |
| CP-578 | Acropora digitifera | Chromoprotein |
| CP-579 | Acropora nobilis | Chromoprotein |
| CP-580 | Acropora hyacinthus | Chromoprotein |
| CP-580 | Acropora hyacinthus | Chromoprotein |
| CP-584 | Acropora nobilis | Chromoprotein |
| CP-584 | Acropora pulchra | Chromoprotein |
| CP-586 | Acropora sp. | Chromoprotein |
| CP-588 | Acropora formosa | Chromoprotein |
| CP-588 | Acropora millepora | Chromoprotein |
| CP-588 | Acropora millepora | Chromoprotein |
| CP-591 | Acropora nobilis | Chromoprotein |
| P-400 | Acropora horrida | CFP |
| P-409 | Acropora nastua | CFP |
| P-445 | Acropora aspera | CFP |
| P-476 | Acropora aspera (orange band I)* | CFP |
| P-478 | Acropora aspera (orange band II)* | CFP |
| P-480 | Acropora aspera (green band)* | CFP |
| P-480 | Acropora tenuis (green band)* | CFP |
| P-482 | Acropora nastua | CFP |
| P-482 | Acropora secale (green band)* | CFP |
| P-483 | Acropora nastua (green band)* | CFP |
| P-483 | Acropora pulchra | CFP |
| P-484 | Acropora millepora | CFP |
| P-484 | Acropora secale | CFP |
| P-485 | Acropora horrida | CFP |
| P-485 | Acropora tenuis | CFP |
| P-486 | Acropora nobilis | CFP |
| P-487 | Acropora cervicornis | CFP |
| P-489 | Acropora millepora | CFP |
| P-490 | Acropora aspera | CFP |
| P-490 | Acropora digitifera | CFP |
| P-490 | Acropora millepora | CFP |
| P-490 | Acropora nobilis | CFP |
| P-495 | Acropora digitifera | CFP |
| P-495 | Acropora nobilis | CFP |
| P-495 | Acropora sp. | CFP |
| P-497 | Acropora millepora | CFP |
| P-500 | Acropora aspera | GFP |
| P-502 | Acropora aculeus | GFP |
| P-504 | Acropora millepora | GFP |
| P-511 | Acropora nobilis | GFP |
| P-512 | Acropora millepora | GFP |
| P-512 | Acropora millepora | GFP |
| P-513 | Acropora aculeus | GFP |
| P-514 | Acropora aspera | GFP |
| P-515 | Acropora eurystoma (now A. tenuis) | GFP |
| P-515 | Acropora tenuis (Alieva listed as A. eurystoma) | GFP |
| P-517 | Acropora tenuis | GFP |
| P-517 | Acropora yongei | GFP |
| P-518 | Acropora cytheria @ Waikiki Aquarium | GFP |
| P-518 | Acropora digitifera | GFP |
| P-590 | Acropora digitifera | DsRed |
| P-593 | Acropora millepora | DsRed |
| P-597 | Acropora millepora | DsRed |
| P-625 | Acropora horrida | DsRed |
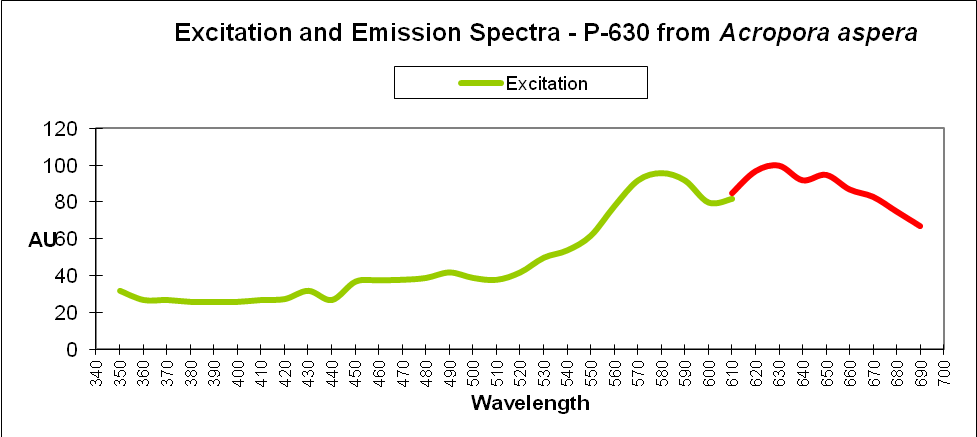
| P-630 | Acropora aspera | DsRed |
Step 3: Expand Available Information to Make It Easy to Use
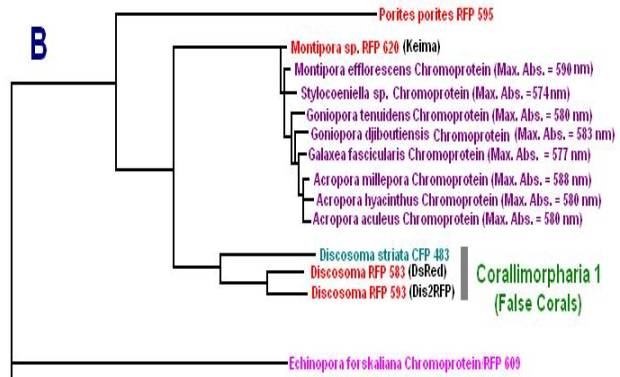
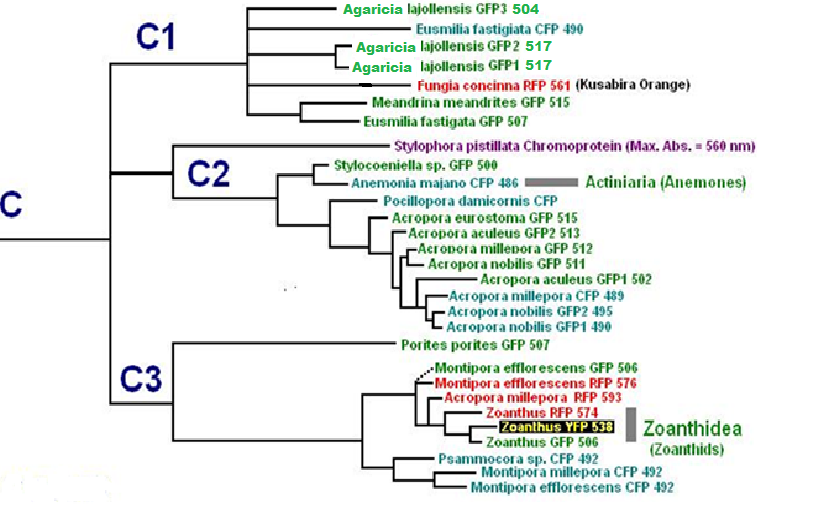
Now, we’ll add additional information to our list. Researchers have analyzed protein structures and then classified coral proteins according to their common ancestors. A group of individual proteins sharing a common origin is called a clade. In Cnidaria, there are 7 colorful clades (A, B, C1, C2, C3, D and Chromo-red). Acropora species are currently known to contain clades B, C2, and C3. See Figures below for a visualization of the relationship of individuals within clades. The phylogenetic trees below are from Alieva et al. (2008). I have modified the list by adding additional information garnered from various sources.
Figure 10. Clade B includes non-fluorescent chromoproteins found in many Acropora species.
Figure 11. Pigment Clades C2 and C3 are of interest when studying Acropora coloration. Note how closely the Acropora proteins are related.
Step 4: Add Clade Information To Our List
Note all non-fluorescent Acropora chromoproteins are in Clade B.
| Emission/Abs | Host | Type | Clade |
|---|---|---|---|
| CP-588 | Acropora formosa | Chromoprotein | B |
| CP-579 | Acropora nobilis | Chromoprotein | B |
| CP-584 | Acropora nobilis | Chromoprotein | B |
| CP-591 | Acropora nobilis | Chromoprotein | B |
| CP-580 | Acropora hyacinthus | Chromoprotein | B |
| CP-580 | Acropora hyacinthus | Chromoprotein | B |
| CP-578 | Acropora digitifera | Chromoprotein | B |
| CP-578 | Acropora digitifera | Chromoprotein | B |
| CP-588 | Acropora millepora | Chromoprotein | B |
| CP-588 | Acropora millepora | Chromoprotein | B |
| CP-584 | Acropora pulchra | Chromoprotein | B |
| CP-580 | Acropora aculeus | Chromoprotein | B |
| CP-580 | Acropora aculeus | Chromoprotein | B |
| CP-500 | Acropora sp. | Chromoprotein | B |
| CP-586 | Acropora sp. | Chromoprotein | B |
| P-487 | Acropora cervicornis | CFP | C2 |
| P-486 | Acropora nobilis | CFP | C2 |
| P-490 | Acropora nobilis | CFP | C2 |
| P-495 | Acropora nobilis | CFP | C2 |
| P-511 | Acropora nobilis | GFP | C2 |
| P-517 | Acropora yongei | GFP | C2 |
| P-400 | Acropora horrida | CFP | C2 |
| P-485 | Acropora horrida | CFP | C2 |
| P-518 | Acropora cytheria @ Waikiki Aquarium | GFP | C2 |
| P-490 | Acropora digitifera | CFP | C2 |
| P-495 | Acropora digitifera | CFP | C2 |
| P-518 | Acropora digitifera | GFP | C2 |
| P-484 | Acropora millepora | CFP | C2 |
| P-489 | Acropora millepora | CFP | C2 |
| P-490 | Acropora millepora | CFP | C2 |
| P-497 | Acropora millepora | CFP | C2 |
| P-504 | Acropora millepora | GFP | C2 |
| P-512 | Acropora millepora | GFP | C2 |
| P-512 | Acropora millepora | GFP | C2 |
| P-445 | Acropora aspera | CFP | C2 |
| P-490 | Acropora aspera | CFP | C2 |
| P-500 | Acropora aspera | GFP | C2 |
| P-514 | Acropora aspera | GFP | C2 |
| P-480 | Acropora aspera (green band)* | CFP | C2 |
| P-476 | Acropora aspera (orange band I)* | CFP | C2 |
| P-478 | Acropora aspera (orange band II)* | CFP | C2 |
| P-483 | Acropora pulchra | CFP | C2 |
| P-515 | Acropora eurystoma (now A. tenuis) | GFP | C2 |
| P-485 | Acropora tenuis | CFP | C2 |
| P-517 | Acropora tenuis | GFP | C2 |
| P-515 | Acropora tenuis (Alieva listed as A. eurystoma) | GFP | C2 |
| P-480 | Acropora tenuis (green band)* | CFP | C2 |
| P-484 | Acropora secale | CFP | C2 |
| P-482 | Acropora secale (green band)* | CFP | C2 |
| P-409 | Acropora nastua | CFP | C2 |
| P-482 | Acropora nastua | CFP | C2 |
| P-483 | Acropora nastua (green band)* | CFP | C2 |
| P-495 | Acropora sp. | CFP | C2 |
| P-625 | Acropora horrida | DsRed | C3 |
| P-590 | Acropora digitifera | DsRed | C3 |
| P-593 | Acropora millepora | DsRed | C3 |
| P-597 | Acropora millepora | DsRed | C3 |
| P-630 | Acropora aspera | DsRed | C3 |
| P-502 | Acropora aculeus | GFP | C3 |
| P-513 | Acropora aculeus | GFP | C3 |
Step 5: Classification
Veron (2000) has classified Acropora species into 38 groups. Appropriate information is now added to our list.
| Emission/Abs | Host | Type | Clade | Group |
|---|---|---|---|---|
| P-487 | Acropora cervicornis | CFP | C2 | 6 |
| CP-588 | Acropora formosa | Chromoprotein | B | 6 |
| CP-579 | Acropora nobilis | Chromoprotein | B | 7 |
| CP-584 | Acropora nobilis | Chromoprotein | B | 7 |
| CP-591 | Acropora nobilis | Chromoprotein | B | 7 |
| P-486 | Acropora nobilis | CFP | C2 | 7 |
| P-490 | Acropora nobilis | CFP | C2 | 7 |
| P-495 | Acropora nobilis | CFP | C2 | 7 |
| P-511 | Acropora nobilis | GFP | C2 | 7 |
| P-517 | Acropora yongei | GFP | C2 | 13 |
| P-400 | Acropora horrida | CFP | C2 | 14 |
| P-485 | Acropora horrida | CFP | C2 | 14 |
| P-625 | Acropora horrida | DsRed | C3 | 14 |
| P-518 | Acropora cytheria @ Waikiki Aquarium | GFP | C2 | 19 |
| CP-580 | Acropora hyacinthus | Chromoprotein | B | 19 |
| CP-580 | Acropora hyacinthus | Chromoprotein | B | 19 |
| CP-578 | Acropora digitifera | Chromoprotein | B | 22 |
| CP-578 | Acropora digitifera | Chromoprotein | B | 22 |
| P-490 | Acropora digitifera | CFP | C2 | 22 |
| P-495 | Acropora digitifera | CFP | C2 | 22 |
| P-518 | Acropora digitifera | GFP | C2 | 22 |
| P-590 | Acropora digitifera | DsRed | C3 | 22 |
| CP-588 | Acropora millepora | Chromoprotein | B | 25 |
| CP-588 | Acropora millepora | Chromoprotein | B | 25 |
| P-484 | Acropora millepora | CFP | C2 | 25 |
| P-489 | Acropora millepora | CFP | C2 | 25 |
| P-490 | Acropora millepora | CFP | C2 | 25 |
| P-497 | Acropora millepora | CFP | C2 | 25 |
| P-504 | Acropora millepora | GFP | C2 | 25 |
| P-512 | Acropora millepora | GFP | C2 | 25 |
| P-512 | Acropora millepora | GFP | C2 | 25 |
| P-593 | Acropora millepora | DsRed | C3 | 25 |
| P-597 | Acropora millepora | DsRed | C3 | 25 |
| P-445 | Acropora aspera | CFP | C2 | 26 |
| P-490 | Acropora aspera | CFP | C2 | 26 |
| P-500 | Acropora aspera | GFP | C2 | 26 |
| P-514 | Acropora aspera | GFP | C2 | 26 |
| P-630 | Acropora aspera | DsRed | C3 | 26 |
| P-480 | Acropora aspera (green band)* | CFP | C2 | 26 |
| P-476 | Acropora aspera (orange band I)* | CFP | C2 | 26 |
| P-478 | Acropora aspera (orange band II)* | CFP | C2 | 26 |
| CP-584 | Acropora pulchra | Chromoprotein | B | 26 |
| P-483 | Acropora pulchra | CFP | C2 | 26 |
| P-502 | Acropora aculeus | GFP | C3 | 28 |
| P-513 | Acropora aculeus | GFP | C3 | 28 |
| CP-580 | Acropora aculeus | Chromoprotein | B | 28 |
| CP-580 | Acropora aculeus | Chromoprotein | B | 28 |
| P-515 | Acropora eurystoma (now A. tenuis) | GFP | C2 | 29 |
| P-485 | Acropora tenuis | CFP | C2 | 29 |
| P-517 | Acropora tenuis | GFP | C2 | 29 |
| P-515 | Acropora tenuis (Alieva listed as A. eurystoma) | GFP | C2 | 29 |
| P-480 | Acropora tenuis (green band)* | CFP | C2 | 29 |
| P-484 | Acropora secale | CFP | C2 | 33 |
| P-482 | Acropora secale (green band)* | CFP | C2 | 33 |
| P-409 | Acropora nastua | CFP | C2 | 34 |
| P-482 | Acropora nastua | CFP | C2 | 34 |
| P-483 | Acropora nastua (green band)* | CFP | C2 | 34 |
| CP-500 | Acropora sp. | Chromoprotein | B | ? |
| CP-586 | Acropora sp. | Chromoprotein | B | ? |
| P-495 | Acropora sp. | CFP | C2 | ? |
Step 6: Add comments
We have now sorted Acropora species according to color, clade, and group. There is a last step – add comments and sort by species in order to facilitate use.
| Emission/Abs | Host | Type | Clade | Group | Comments |
|---|---|---|---|---|---|
| P-502 | Acropora aculeus | GFP | C3 | 28 | 2 spp., distinct from other groups |
| P-513 | Acropora aculeus | GFP | C3 | 28 | 2 spp., distinct from other groups |
| CP-580 | Acropora aculeus | Chromoprotein | B | 28 | 2 spp., distinct from other groups |
| CP-580 | Acropora aculeus | Chromoprotein | B | 28 | 2 spp., distinct from other groups |
| P-445 | Acropora aspera | CFP | C2 | 26 | Distinct unit, but related to 25 |
| P-490 | Acropora aspera | CFP | C2 | 26 | Distinct unit, but related to 25 |
| P-500 | Acropora aspera | GFP | C2 | 26 | Distinct unit, but related to 25 |
| P-514 | Acropora aspera | GFP | C2 | 26 | Distinct unit, but related to 25 |
| P-630 | Acropora aspera | DsRed | C3 | 26 | Distinct unit, but related to 25 |
| P-480 | Acropora aspera (green band)* | CFP | C2 | 26 | Distinct unit, but related to 25 |
| P-476 | Acropora aspera (orange band I)* | CFP | C2 | 26 | Distinct unit, but related to 25 |
| P-478 | Acropora aspera (orange band II)* | CFP | C2 | 26 | Distinct unit, but related to 25 |
| P-487 | Acropora cervicornis | CFP | C2 | 6 | * |
| P-518 | Acropora cytheria @ Waikiki Aquarium | GFP | C2 | 19 | * |
| CP-578 | Acropora digitifera | Chromoprotein | B | 22 | * |
| CP-578 | Acropora digitifera | Chromoprotein | B | 22 | * |
| P-490 | Acropora digitifera | CFP | C2 | 22 | * |
| P-495 | Acropora digitifera | CFP | C2 | 22 | * |
| P-518 | Acropora digitifera | GFP | C2 | 22 | * |
| P-590 | Acropora digitifera | DsRed | C3 | 22 | * |
| P-515 | Acropora eurystoma (now A. tenuis) | GFP | C2 | 29 | * |
| CP-588 | Acropora formosa | Chromoprotein | B | 6 | * |
| P-400 | Acropora horrida | CFP | C2 | 14 | * |
| P-485 | Acropora horrida | CFP | C2 | 14 | * |
| P-625 | Acropora horrida | DsRed | C3 | 14 | * |
| CP-580 | Acropora hyacinthus | Chromoprotein | B | 19 | * |
| CP-580 | Acropora hyacinthus | Chromoprotein | B | 19 | * |
| CP-588 | Acropora millepora | Chromoprotein | B | 25 | Distinct unit, but related to 26 |
| CP-588 | Acropora millepora | Chromoprotein | B | 25 | Distinct unit, but related to 26 |
| P-484 | Acropora millepora | CFP | C2 | 25 | Distinct unit, but related to 26 |
| P-489 | Acropora millepora | CFP | C2 | 25 | Distinct unit, but related to 26 |
| P-490 | Acropora millepora | CFP | C2 | 25 | Distinct unit, but related to 26 |
| P-497 | Acropora millepora | CFP | C2 | 25 | Distinct unit, but related to 26 |
| P-504 | Acropora millepora | GFP | C2 | 25 | Distinct unit, but related to 26 |
| P-512 | Acropora millepora | GFP | C2 | 25 | Distinct unit, but related to 26 |
| P-512 | Acropora millepora | GFP | C2 | 25 | Distinct unit, but related to 26 |
| P-593 | Acropora millepora | DsRed | C3 | 25 | Distinct unit, but related to 26 |
| P-597 | Acropora millepora | DsRed | C3 | 25 | Distinct unit, but related to 26 |
| P-409 | Acropora nastua | CFP | C2 | 34 | * |
| P-482 | Acropora nastua | CFP | C2 | 34 | * |
| P-483 | Acropora nastua (green band)* | CFP | C2 | 34 | * |
| CP-579 | Acropora nobilis | Chromoprotein | B | 7 | * |
| CP-584 | Acropora nobilis | Chromoprotein | B | 7 | * |
| CP-591 | Acropora nobilis | Chromoprotein | B | 7 | * |
| P-486 | Acropora nobilis | CFP | C2 | 7 | * |
| P-490 | Acropora nobilis | CFP | C2 | 7 | * |
| P-495 | Acropora nobilis | CFP | C2 | 7 | * |
| P-511 | Acropora nobilis | GFP | C2 | 7 | * |
| CP-584 | Acropora pulchra | Chromoprotein | B | 26 | Distinct unit, but related to 25 |
| P-483 | Acropora pulchra | CFP | C2 | 26 | Distinct unit, but related to 25 |
| P-484 | Acropora secale | CFP | C2 | 33 | * |
| P-482 | Acropora secale (green band)* | CFP | C2 | 33 | * |
| CP-500 | Acropora sp. | Chromoprotein | B | ? | * |
| CP-586 | Acropora sp. | Chromoprotein | B | ? | * |
| P-495 | Acropora sp. | CFP | C2 | ? | * |
| P-485 | Acropora tenuis | CFP | C2 | 29 | * |
| P-517 | Acropora tenuis | GFP | C2 | 29 | * |
| P-515 | Acropora tenuis (Alieva listed as A. eurystoma) | GFP | C2 | 29 | * |
| P-480 | Acropora tenuis (green band)* | CFP | C2 | 29 | * |
| P-517 | Acropora yongei | GFP | C2 | 13 | * |
We now have a small database for comparative purposes and hence make some educated guesses about the relationship of colors among Acropora species and their functions (I’ve consolidated much information already) – this has been relatively easy. This listing is rather small, containing about 60 entries of corals in one genus. The entire database (not shown) contains over 300 entries spanning about 50 genera. There is one other database I’ve compiled that is of use when examining the effects of light on corals, and this list is of different sorts of symbiotic dinoflagellates (Symbiodinium species, or ‘zooxanthellae’) found in corals. A coral (called the host) may contain one of more types of zooxanthellae (also divided into groups called clades). Note that the groups of zooxanthellae clades might share the same alpha-numeric name with a fluorescent protein clade. This is strictly coincidence and a coral with Clade C1 zooxanthellae can also contain a colorful protein called C1. It is also possible that a coral with an alga belonging to Clade D can contain a fluorescent color of Clade B. Whatever the case, the zooxanthellae clade list consists of about 2,000 entries and is much too long to re-post here. See here for further information about coral zooxanthellae: http://www.advancedaquarist.com/2011/4/aafeature
Step 7: Review literature and compile known information about Acropora colors, possible functions, and other data
This information has recently become available in the last few years. After this quick review, we’ll attempt to synthesize the data and try to make some sense of it.
Acropora yongei (commonly called the Bali Green Slimer by many hobbyists): Veron’s Acropora Group 13. Contains zooxanthellae clade C3 and Green Fluorescent Protein (GFP) with an emission at 517nm (most likely of Protein CladeC2).
Acropora millepora: Veron’s Acropora Group Group 25 & Zooxanthellae Clade C3*. Fluorescent Proteins described to be found in A. millepora are cyan (with emissions at 484, 489, and 497nm, a green fluorescent protein with a fluorescence peak at 512nm, and at least red ones (emission at 593nm and 597nm – all the cyan and green fluorescent proteins are of likely of Clade C2 while red is probably from Clade C3). There is also a blue non-fluorescent protein with a maximum absorbance at 588nm; Protein Clade B).
*According to reports, Acropora millepora most often contains Clade C3 zooxanthellae (a Generalist Clade), but may also possess these clades as well (listed in descending order): C1 (also a Generalist Clade), C2, C4, C3k, and D.
Acropora pulchra is of Veron’s Acropora Group 26 – closely related to those Acropora species of Group 25, such as A. millepora, above) & Zooxanthellae Clade C3. This species is known to contain a fluorescent cyan protein (emission at483nm, and a non-fluorescent protein with a maximum absorbance at 584nm). The cyan protein – based on best available information -is likely of Protein Clade C2, and the non-fluorescent one is Clade B.
We see remarkable similarities: Although we can’t be sure that all these Acropora corals contain the Generalist Zooxanthella Clade C3, it would be a good bet to make. This is an important factor. Zooxanthellae of Clade C3 are not as tolerant of high light as some other zooxanthellae clades, but they are highly adaptable to a wide range of illumination. The best information we currently have says the Protein Clades are B, C2, and C3. If we believe these assumptions are correct, we see trends evolve.
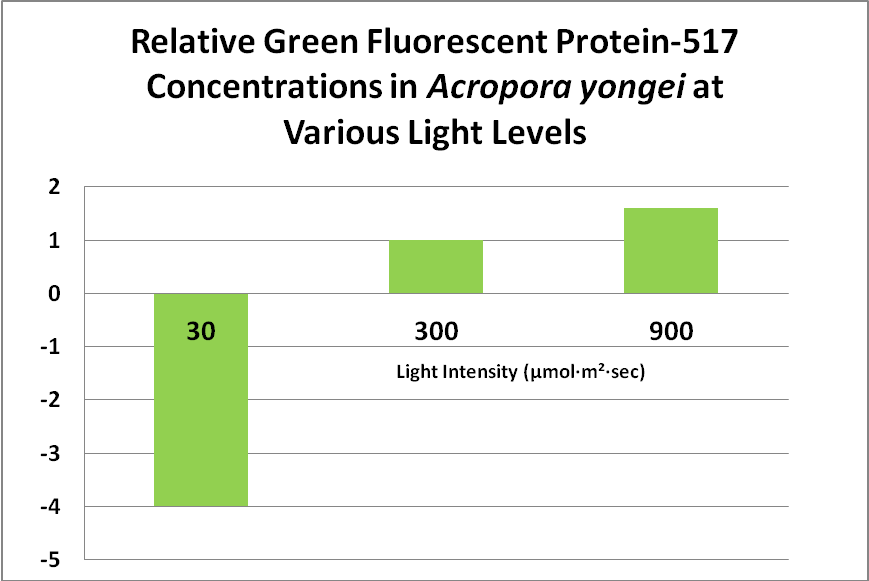
Role of Fluorescent and Non-fluorescent Proteins: Kawaguti (1944) made remarkable observations and believed coral colors were natural sunscreens. Roth et al. (2010) have confirmed this in a tangential sort of way. The latter scientists subjected Acropora yongei (the Bali Slimer Acropora, if you will) fragments to various light intensities using T5 fluorescent lamps. PAR values were categorized as follows: Low Light (30 µmol·m²·sec), Medium Light (300 µmol·m²·sec) and High Light (900 µmol·m²·sec -a remarkable feat considering the light source). They observed the green fluorescence under all lighting conditions. In low light conditions, the density (number) of zooxanthellae fell to very low levels, as did the level of GFP. This is contrary to what many hobbyists believe – that zooxanthellae numbers would increase under low light (causing a ‘browning’ of the coral thus masking the GFP). These researchers believe the loss of zooxanthellae might be in an effort to avoid self-shading of the dinoflagellates, or perhaps due to a reduction in volume of coral host tissues. GFP content increased substantially at 300 µmol·m²·sec, and increased by 1.6X at a light level of 900 µmol·m²·sec. See Figure 13.
Figure 13. This green fluorescent protein (GFP-517) becomes more apparent at higher light levels. After Roth et al., 2010.
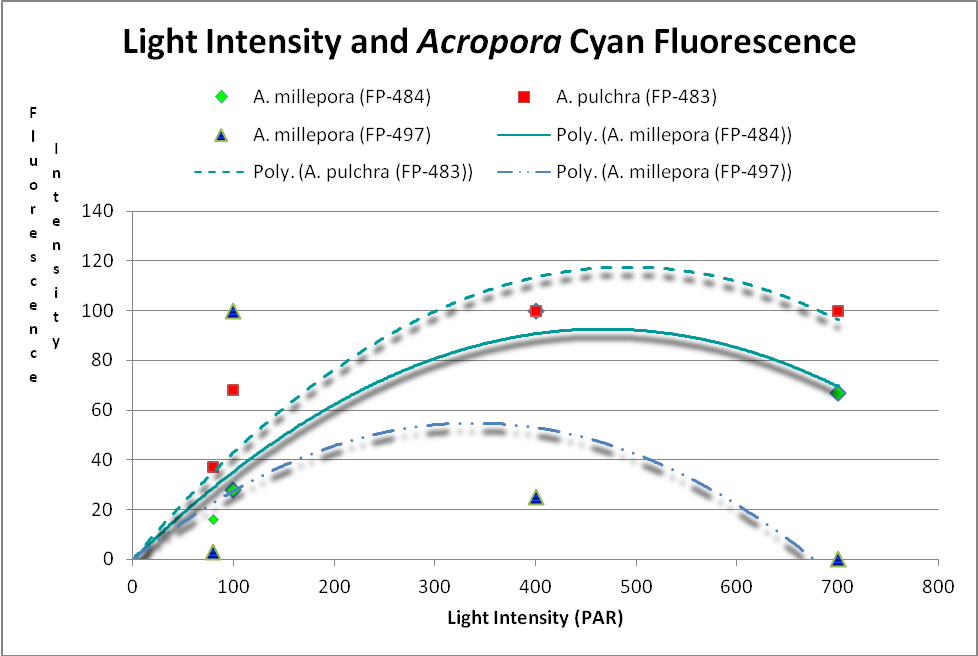
These researchers clearly linked Acropora pigment concentration and fluorescence intensity to light intensity. Other reports reached the same conclusion. D’Angelo et al. (2008; 2011) light intensity played a part in the corals’ productions of fluorescent pigments from Acropora pulchra and A. millepora. Broadband blue light (as opposed to green and red light) was most effective in promoting fluorescence in Acropora pulchra proteins (a cyan protein with emission at 483nm and a chromoprotein with maximum absorbance at 584nm) as well as those fluorescent Acropora millepora pigments (emissions at 484, 497, 512 and 597nm). See Figures 14- 17.
Figure 14. Cyan (blue-green) fluorescent proteins are down-regulated at higher light intensities. After D’Angelo et al., 2011.
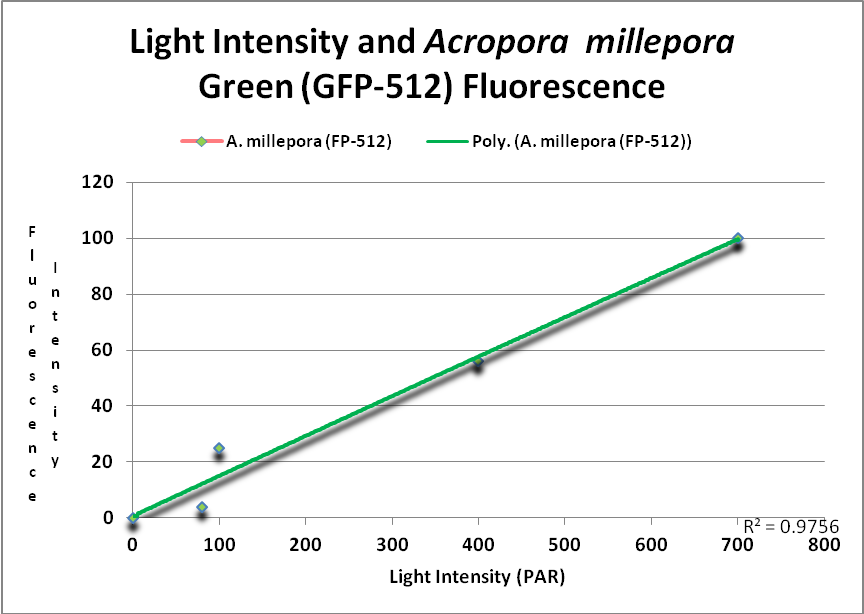
Figure 15. This GFP (green fluorescent protein or GFP-512) is expressed as light levels increase. After D’Angelo et al., 2011.
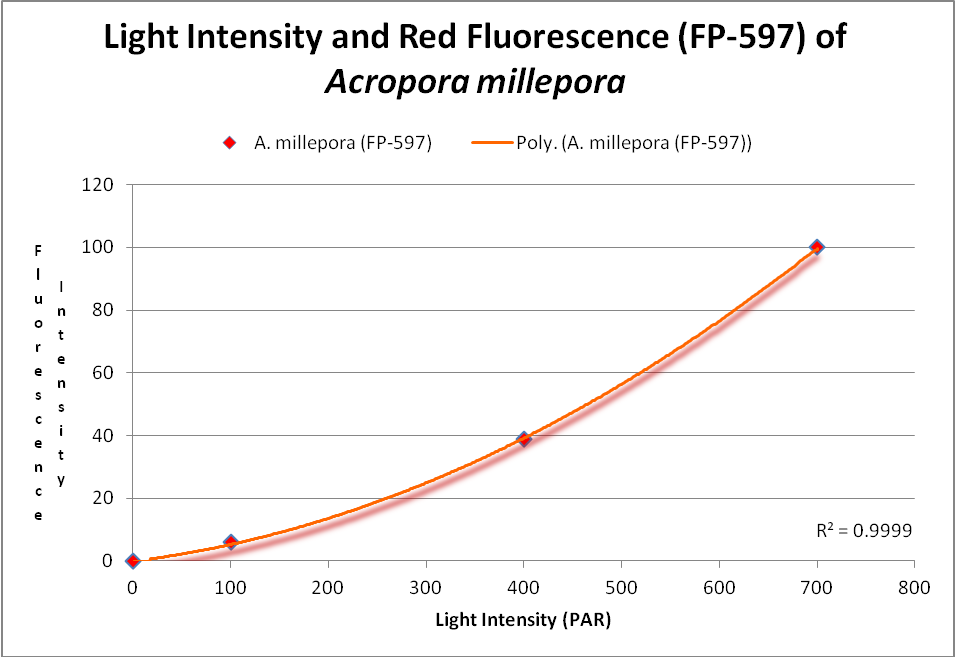
Figure 16. This RFP (red fluorescent protein) increases with light intensity. After D’Angelo et al., 2011.
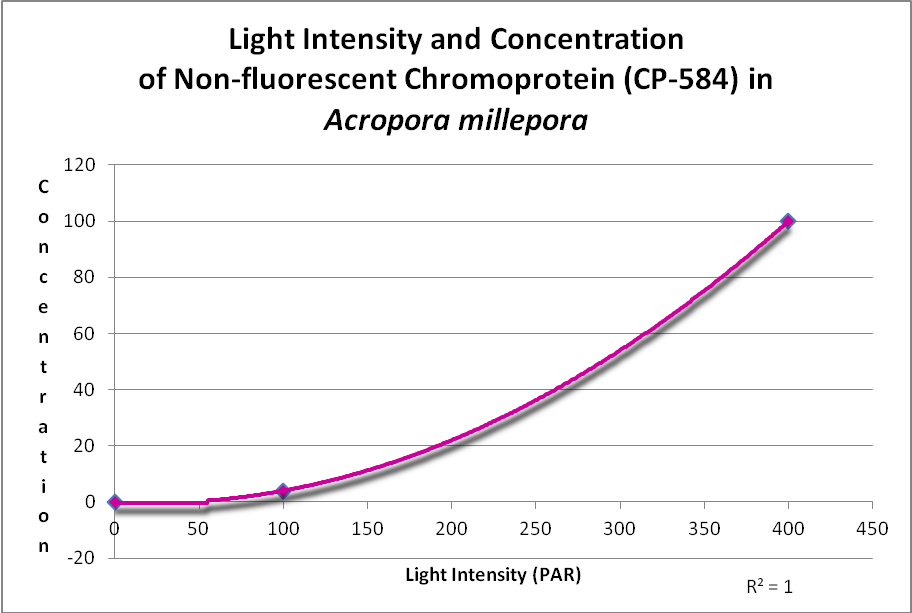
In addition, a non-fluorescent chromoprotein (CP-584) from an Acropora millepora also demonstrated an increase in concentration with increasing light intensities. See Figure 17.
Figure 17. This chromoprotein increases with light intensity. It makes the Acropora appear purple-blue. After D’Angelo et al., 2011.
As aquarists have long observed, there is often an increase in coral coloration when light intensity increases, especially when the light spectrum is skewed towards the blue portion. But why?
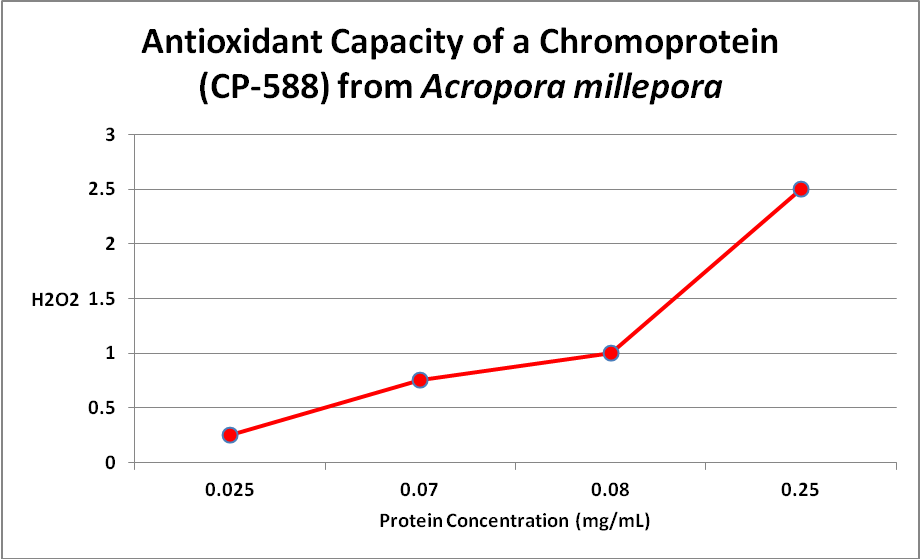
Palmer et al. (2009) examined the relationship of several fluorescent and one non-fluorescent proteins’ concentration and anti-oxidant capacity. Several have speculated about this ability, but this is the first experiment I’m aware of that examines this. Only one Acropora chromoprotein was examined – that of an Acropora millepora -but there was clear evidence that an oxidant (hydrogen peroxide, or H 2O2) was destroyed by the protein. See Figure 18.
Figure 18. More damaging hydrogen peroxide (H2O2) is scavenged (removed) as the amount of non-fluorescent protein increases. The same happens with coral fluorescent proteins. From Palmer et al., 2009.
This is important -H 2O2 is produced by the coral animal but especially so by its zooxanthellae. Hydrogen peroxide is destructive to living tissues and there are metallic enzymes that can ‘detoxify’ it. However, under conditions of high light and/or high temperature stress, the coral animal can produce natural defenses – fluorescent and/or non-fluorescent chromoproteins. Palmer et al. () also examined the antioxidant capacities of fluorescent proteins as well as the non-fluorescent chromoprotein. When the pigments were isolated from the coral and purified, the chromoprotein had the highest antioxidant capacity, followed by cyan, red, and green fluorescent proteins. However, the amount of fluorescent proteins varies within corals and when these researchers looked at in vivo anti-oxidation capacities, cyan FP did not have and hydrogen peroxide scavenging due to low content. On the other hand, GFP has a relatively low ability to deal with oxygen radicals but is found in high concentrations, thus it is able to deal with H 2O2. The red fluorescent protein was quite effective in vivo, but the non-fluorescent choromoprotein was most efficient. The pieces are finally falling into place.
D’Angelo et al. (2012) offer more evidence for the role of corals’ coloration. They hypothesize that the usually non-fluorescent chromoprotein found in fast-growing (or recently damaged) tissues are generated by the coral animal in response to high light levels (where the white coral skeleton is free of zooxanthellae and strongly reflects light, thereby maximizing intensity). These researchers also found growth markers in tips, margins and damaged areas where growth is quickest.
Effects of Temperature
Temperature has a major impact on many biological functions, and the expression of fluorescent proteins within corals is no exception. Down-regulation (lack of production) of RFP-593 in Acropora millepora occurs at temperatures of 32 – 33 (89.6 – 91.4). Some Anthozoan fluorescent proteins mature faster at temperatures 30C (86F; Terskikh et al., 2002.)
Upregulation of a blue chromoprotein (CP-588) in Acropora millepora specimens has been noted during a natural bleaching event on Australia’s Great Barrier Reef when temperature plateaued at 32C (89.6).
Effects of pH
Papina et al. (2002) found no significant variations in the fluorescent signatures of proteins from several Acropora species at pH values ranging from 5 to 8.
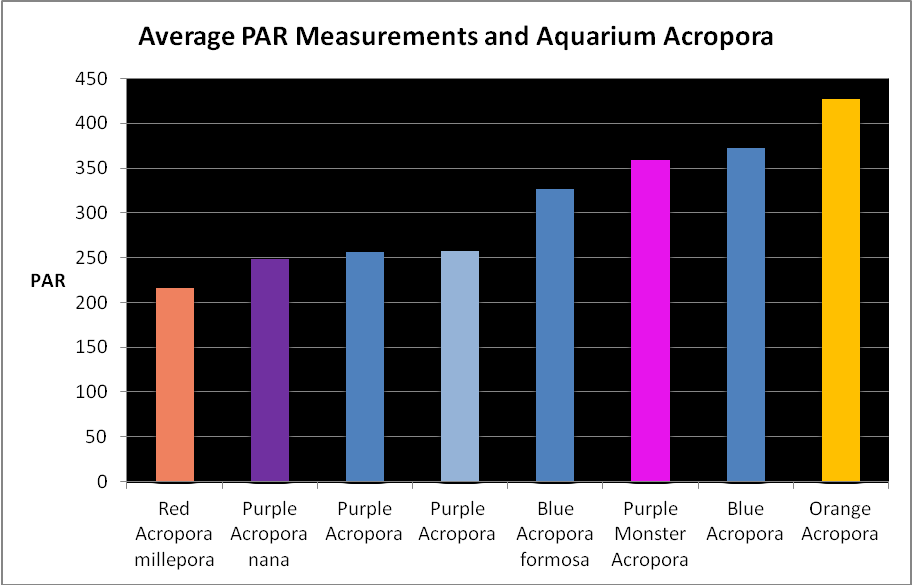
Figure 19. Average light measurements of variously colored aquarium Acropora specimens (species based on best guess). All are based on at least 6 PAR measurements using a Li-Cor quantum meter. Many of these measurements were of Iwasaki 6500K ‘daylight’ metal halide lamps and Very High Output (VHO) actinic and daylight fluorescent lamp combinations (quite popular in the 1990’s). So many miles, so many living rooms…
Light Intensity
Figure 19 shows average light intensities used to maintain variously colored Acropora specimens. These were collected by me over the years while visiting hundreds of hobbyists. They kindly allowed me to take these measurements with a Li-Cor quantum meter.
Light Spectral Quality
Hobbyists have long made anecdotal observations linking coral coloration and blue light. D’Angelo et al. (2011) investigated the effects of broad bandwidths (blue, green, and red) on coral color generation using metal halide lamps and theatrical light filters. Blue light best promoted green and red fluorescence in Acropora millepora specimens. Green light promoted green fluorescence and marginal red fluorescence. Red light did not produce noticeable results. Riddle (2003) tested the effects equal light intensity from red and blue LEDs on a Pocillopora meandrina. See: http://www.advancedaquarist.com/2003/11/aafeature (Synopsis of this article: The blue LED caused the expression of a pink chromoprotein, while the red LED bleached the coral.) Verkhusha and Lukyanov (2004) found infrared radiation would cause DsRed to become green.
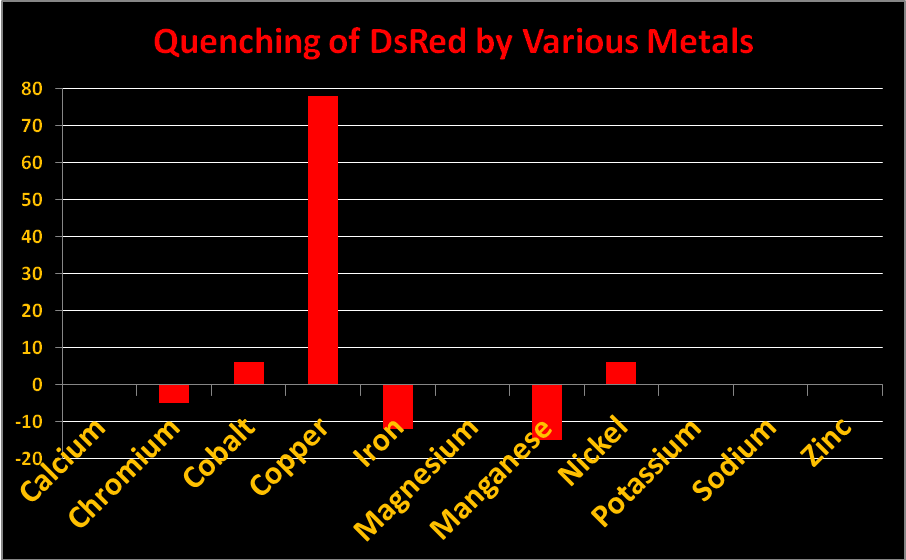
Figure 20. Reduction in fluorescence is called quenching. Some metals quench fluorescence of Acropora DsRed-type pigments while others do not.
Metals
Certain metals can alter the charge of the protein barrel of DsRed pigments thus changing fluorescence intensity. The effect is called ‘quenching’ when fluorescence decreases. In Acropora specimens, orange or red colors are due to DsRed-type proteins. In Figure 20, we see chromium, iron, and manganese will slightly increase fluorescence while copper, cobalt, and manganese will quench it. Other metals (shown) had no effect. Resist the temptation to haphazardly dump any metal into your aquarium!
In Closing
Fluorescent and non-fluorescent proteins are found in many Acropora specimens (those genetically predisposed to express them). The specimens cover a wide range of Acropora groups (as described by Veron) ranging from tabletops to corymbose bushes – no relationship between coral morphology and coloration was established.
Description of GFP-like protein clades allows us to visualize their relationships. Thus we can make assumptions and make generalizations as to how these proteins would react to various stimuli.
We seem to finally have an answer for several of the roles of coral coloration. The ability of the coral to produce colored antioxidants (in the form of hydrogen peroxide scavenging fluorescent/non-fluorescent proteins) has ramifications – a lighter colored coral absorbs less heat than a brown coral (Fabricius, 2006). Some of the absorbed light might be fluoresced away from the portions of the spectrum absorbed by zooxanthellae for photosynthesis. Light reflected lessens that absorbed by photosynthetic pigments (Mazel, personal communication). While the contribution of each of these factors might be in and of itself small, the overall effect might be significant.
Perhaps of more importance to the hobbyist is the realization that we can manipulate the lighting environment in order to create and/or maintain color coloration. It has long been my contention that we should concentrate on minimum light levels to achieve the desired results. The thought of using 1,000 watt metal halide lamps over shallow aquaria now seems so wasteful (worse yet was the thought to use the Fusion Sulfur Lamp capable of producing a PAR value in excess of 20,000 µmol·m²·sec – one magnitude brighter than the sun at noon on a cloudless day).
Of the Acropora species examined here, most likely contain ‘generalist’ zooxanthellae clades C1 or C3. These clades are highly adaptable to light intensity if allowed to acclimate slowly (weeks) and are found in a number of coral taxa. Some Acropora might contain zooxanthellae clade C2, or one of the C3 sub-clades (clade C2 is found almost exclusively in Acropora specimens. The Great Barrier Reef and Taiwan are listed as collection sites. Another paper reports Pocillopora specimens to contain C3. Clade C3i has been found only in about 10 Acropora species at relatively shallow depths of 1-10m (~3 feet to 33 feet). Clade C3i is believed to have descended from zooxanthellae Clade C1. Atlantic Acropora species (A. cervicornis and A. palmata) contain zooxanthellae of Clade A.
What does it tell us about the amount of light required by the captive dinoflagellates when their host generates protective colorful proteins in order to protect itself from harmful by-products of photosynthesis (such as hydrogen peroxide)? Acropora specimens, as a general rule, are thought of as ‘light-loving’ by many hobbyists. Does this concept hold water when the coral produces protective pigments at light levels of, say, 200 µmol·m²·sec?
Cyan FPs’ contents peak at less than 500 µmol·m²·sec, and then decrease (at least in CFP- 483, 484, and 497). Researchers have determined that this is due, in some cases, to down-regulation – it is in the coral’s genes to do so. We cannot be sure that all cyan proteins react in the same manner, and another possibility exists. The double peak excitation and emission shown in Figure 21 suggests photoconversion is possible (possibly a conversion of cyan protein to one that fluoresces green). We know the chromophores of at least some cyan and green fluorescent proteins are similar – it could be this excitation/emission is the result of an incomplete separation of cyan and green pigments during testing through electrophoresis (SDS-PAGE or sodium dodecyl sulfate polyacrylamide gel electrophoresis to be exact).
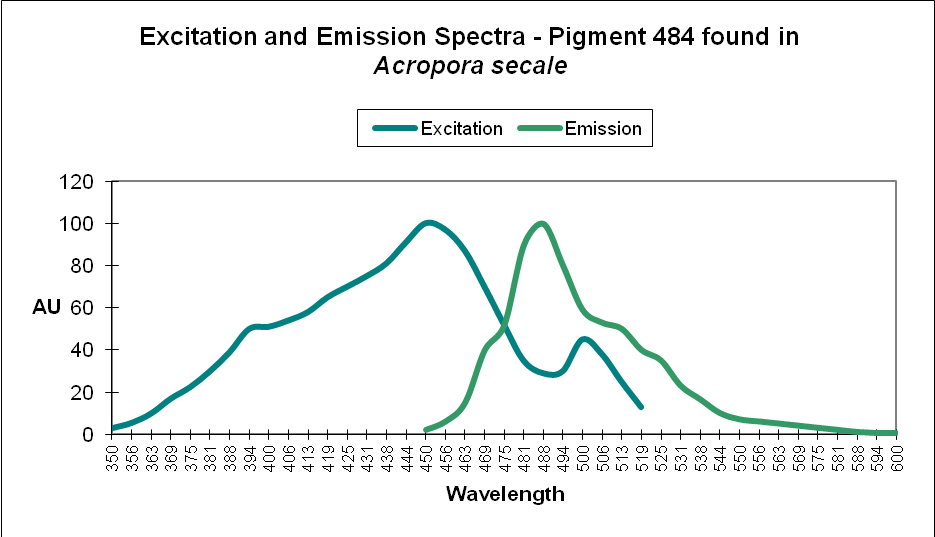
Figure 21. The double peaks seen in the excitation and emissions of this Acropora cyan protein suggests photoconversion is possible. Or is this an artifact of testing? After Papina et al., 2002.
At least two green fluorescent proteins (GFP 512 and 517) will continue to increase at light levels higher than normally seen in aquaria – 700 and 900 µmol·m²·sec, repectively.
Red fluorescence (in Acropora millepora FP-597) will continue to increase at high light intensity (700 µmol·m²·sec). However, ‘greening’ has been noted with some red proteins (mostly those from corals living a depths greater than 40 meters , or 130 feet – in other words, not many). Greening occurs when the red protein is selectively destroyed (photobleached, perhaps by infrared radiation) making the green fluorescence more apparent.
A non-fluorescent chromoprotein (CP-584) from Acropora millepora will also increase as light intensity increases. Chromoproteins have the highest per unit antioxidant capacity of all GFP-like proteins and have been shown to be produced experimentally by the coral at light levels up to 700 µmol·m²·sec (and up to about 1,000 µmol·m²·sec in aquaria).
Temperature excursions (particularly those to 32C and above) can cause color shifts even if the event is short-lived.
Yet, light alone is not enough to maintain coloration. Water movement of velocity sufficient to keep the coral cleansed of sediments is essential. Proper temperature and water chemistry cannot be overlooked. In short, it will take some dedication to maintain an aquarium with glowing Acropora specimens. However, there is no denying that light can be the deciding factor when all other conditions are correct.
There are some factors beyond practical control of the hobbyist, such as the chemical content of artificial seawater where an impurity (such as a metal) might be high (or low) in a batch of sea salts.
When all factors are correct, Acropora corals will as much as 15cm (6 inches) in a year. Their color is a sign of good conditions, where the coral maintains its health in an artificial environment.
Note: I suspect most hobbyists use an Apogee PAR meter to make measurements. These work fine when measuring sunlight and metal halide lamps but they are challenged when used to estimate light output of LEDs. Opinions vary on how to correct the Apogee’s readings. I’ll offer results of some rather tedious testing and suggest correction factors as well as lux-to-PAR conversions. We’ll also examine effects of light on those corals containing Clade D proteins. But those are other stories. More soon.
References
The following 7 references are cited in this article, in addition to the 158 listed below.
- D’Angelo, C., E. Smith, F. Oswald, J. Burt, D. Tchernov, and J. Wiedenmann, 2012. Locally accelerated growth is part of the innate immune response and local repair mechanisms in reef-building corals as detected by green fluorescent protein (GFP)-like pigments. Coral Reefs, 31:1045-1056.
- Palmer, C., M. Roth, and R. Gates, 2009. Red fluorescent protein responsible for pigmentation in trematode-infected Porites compressa tissues. Biol. Bul.. 216: 68-74.
- Palmer, C., C. Modi, and L. Mydiarz, 2009. Coral fluorescent proteins as antioxidants. PlosOne.
- Smith-Keune, C. and S. Dove, 2008. Gene expression of a green fluorescent protein homolog as a host-specific biomarker of heat stress within a reef-building coral. Mar. Biotechnol., 10,2: 166-180
- Roth, M., M. Latz, R. Goericke, and D. Deheyn, 2010. Green fluorescent protein regulation in the coral Acropora yongei during photoacclimation. J. Exp. Biol., 213:3644-3655.
- D’Angelo, C. and J. Wiedenmann, 2011. Blue light and its importance for the colors of stony corals. Coral, 8,6: 64-76.
- Seneca, F., S. Forêt, E. Ball, C. Smith-Keune, D. Miller, and M. van Oppen, 2010. Patterns of gene expression in a scleractinian coral undergoing natural bleaching. Mar. Biotechnol., 12: 594-604
Additional Resources
- Ando, R., H. Hama, M. Yamamoto-Hino, H. Mizuno, and A. Miyawaki, 2002. An optical marker based on the UV-induced green-to-red photoconversion of a fluorescent protein. Proc. Natl. Acad. Sci. USA, 99(20):12651-12656.
- Ando, R., H. Mizuno and A. Miyawaki, 2004. Regulated fast nucleocytoplasmic shuttling observed by reversible protein highlighting. Science, 306:1370-1373.
- Andresen, M., M. Wahl, A. Stiel, F. Gräter, L. Schäfer, S. Trowitzsch, G. Weber, C. Eggeling, H. Grubmüller, S. Hell and S. Jakobs, 205. Structure and mechanism of the reversible photoswitch of a fluorescent protein. Proc. Natl. Acad. Sci. USA, 102, 37:13070-13074.
- Apprill, A., 2003. Spectral characteristics and genetic expression of green fluorescent proteins in Hawaiian corals. In: Molecular Biology of Corals: Results of 2002 Edwin W. Pauley Summer Program in Marine Biology, E. Cox and T. Lewis, eds. University of Hawaii HIMB Technical Report No. 43:6-13.
- Baird, G., D. Zacharias and R. Tsien, 2000. Biochemistry, mutagenesis, and oligomerization of DsRed, a red fluorescent protein from coral. Proc. Natl. Acad. Sci. USA, 97(22):11984-11989.
- Bandaranayake, W., 1998. Mycosporines: Are they nature’s sunscreens? National Product Review, 1998. 159-172.
- Bingman, C., 1995. Green-fluorescent protein: a model for coral host fluorescent proteins? Aquarium Frontiers, 2(3): 6-9.
- Bingman, C., 1999. Biochemistry of Aquaria: Coral Fluorescence – An Update.http://www.reefs.org/library/aquarium_frontiers/index.html
- Blundell, A., 2005. Lateral Lines: The Seen and Unseen World of Coral Fluorescence.http://www.advancedaquarist.com/2005/2/lines/
- Bulina, M., D. Chudakov, N. Mudrik, and K. Lukyanov, 2002. Interconversion of Anthozoa GFP-like fluorescent and non-fluorescent proteins by mutagenesis. BMC Biochem., 24; 3(1):7.
- Bulina, M., K. Lukyanov, I. Yampolsky, D. Chudakov, D. Staroverov, A. Shcheglov, N. Gurskaya and S. Lukyanov, 2004. New class of blue animal pigments based on frizzled and kringle protein domains. J. Biol. Chem., 279(42):43367-43370.
- Burr, A., P. Hunt, D. Wagar, S. Dewilde, M. Blaxter, J. Vanfleteren and L. Moens, 2000. A hemoglobin with an optical function. J. Biol. Chem., 275(7): 4810-4815.
- Calfo, A., 2005. Magnificent fluorescence! Aquaristic perspectives. http://reefkeeping.com/issues/2005-11/ac/index.php
- Chattoraj, M., B. King, G. Bublitz and S. Boxer, 1996. Ultra-fast excited state dynamics in green fluorescent protein: Multiple states and proton transfer. Proc. Natl. Acad. Sci. USA, 93: 8362-8367.
- Chudakov, D., A. Feofanov, N. Mudrik, S. Lukyanov and K. Lukyanov, 2003. Chromophore environment provides clue to “kindling fluorescent protein” riddle. J. Biol. Chem., 278(9):7215-7219.
- Chudakov, D., V. Belousov, A. Zararisky, V. Novoselov, D. Staroverov, D. Zorov, S. Lukyanov and K. Lukyanov, 2003. Kindling fluorescent proteins for precise in vivo photolabeling. Nat. Biotechnol. 21(2): 191-194.
- Cotlet, M., J. Hofkens, S. Habuchi, G. Dirix, M. Van Guyse, J. Michiels, J. Vanderleyden and F. De Schryver, 2001. Identification of different emitting species in the red fluorescent protein DsRed by means of ensemble and single-molecule spectroscopy. Proc. Natl. Acad. Sci. USA, 98(25):14398-14403.
- Cox, G. and A. Salih, 2005. Fluorescent lifetime imaging of symbionts and fluorescent proteins in reef corals. In: Multiphoton Microscopy in the Biomedical Sciences V, edited by A. Periasami and Peter So. Proc. SPIE, 5700:162-170.
- Credabel, J., 2006. Notes from the trenches: Coral fusion and grafting. http://reefkeeping.com/issues/2006-02/nftt/index.php
- D’Elia, C., S. Domotor and K. Webb, 1983. Nutrient uptake kinetics of freshly isolated zooxanthellae. Mar. Biol., 75:157-167.
- Delbeek, J. and J. Sprung, 1994. The Reef Aquarium: A Comprehensive Guide to the Identification and Care of Tropical Marine Invertebrates. Vol. 1. Ricordea Publishing, Coconut Grove, Fl. 544 pp.
- Delbeek, J. and J. Sprung, 2005. The Reef Aquarium: Science, Art and Technology. Vol. 3. Ricordea Publishing, Coconut Grove, Fl. 679 pp.
- Delbeek, J.C., 2003. Media Review, Advanced Aquarist Online Magazine, June. www.advancedaquarist.com/issues/june2003/media.htm
- Dittrich, P., S. Schäfer and P. Schwille, 2005. Characterization of the photoconversion on reaction of the fluorescent protein Kaede on the single-molecule level. Biophys. J., 89:3446-3455.
- Dove, S., M. Takabayashi and O. Hoegh-Guldberg, 1995. Isolation and partial characterization of the pink and blue pigments of Pocilloporid and Acroporid corals. Biol. Bull., 189:288-297.
- Dove, S., O. Hoegh-Guldberg and S. Ranganathan, 2001. Major color patterns of reef-building corals are due to a family of GFP-like proteins. Coral Reefs 19: 197-204.
- Dove, S., 2004. Scleractinian corals with photoprotective host pigments are hypersensitive to thermal bleaching. Mar. Ecol. Prog. Ser., 272: 99-116.
- Dove, S., J. Oritz, S. Enriquez, M. Fine, P. Fisher, R. Iglesias-Prieto, D. Thornhill and O. Hoegh-Guldberg, 2006. Response of holosymbiont pigments from the scleractinian coral Montipora monasteriata to short-term heat stress. Limnol. Oceanogr., 51(2): 1149-1158.
- Elowitz, M., M. Surette, P. Wolf, J. Stock and S. Leibler, 1997. Photoactivation turns green fluorescent protein red. Curr. Biol., 7(10):809-812.
- Fabricius, F., 2006. Effects of irradiance, flow and colony pigmentation on the temperature microenvironment around corals: Implications for coral bleaching. Limnol. Oceanogr., 51(1): 30-37.
- Finet, B. and F. Lesage, 2005. Colors by the thousands – Light, Colors and Corals, Part 1.
- Advanced Aquarist Online, December 2005. http://www.advancedaquarist.com/2005/12/aafeature2
- Fitt, W.K., T.A.V. Rees and D. Yellowlees, 1995. Relationship between pH and the availability of dissolved inorganic nitrogen in the zooxanthella-giant clam symbiosis. Limnol. Oceanogr., 40(5): 976-982.
- Fox, D.L. and D.W. Wilkie, 1970. Somatic and skeletally fixed carotenoids of the purple hydrocoral, Allopora californica. Comp. Biochem. Physiol., 36:49-60.
- Fox, D.L., 1972. Pigmented calcareous skeletons of some corals. Comp. Biochem. Physiol., 43B:919-927.
- Fux, E. and C. Mazel, Unpublished. An experimental method to separate the fluorescence and reflectance components of the spectral signatures of corals.
- Fux, E. and C. Mazel, 1999. Unmixing coral fluorescence emission spectra and predicting new spectra under different excitation conditions. Applied Optics. 38, 3: 486-494.
- Garcia-Parajo, M., M. Koopman, E. van Dijk, V. Subramaniam, and N.F. van Hulst, 2001. The nature of fluorescence emission in the red fluorescent protein DsRed, revealed by single-molecule detection. Proc. Natl. Acad. Sci. USA, 98(25):14392-14397.
- Gentien, P., 1981. Fluorescent metabolites in coral reefs off Townsville, Queensland. Aust. J. Mar. Freshwater Res., 32: 975-980.
- Gilmore, A., A. Larkum, A. Salih, S. Itoh, Y. Shibata, C. Bena, H. Yamasaki, M. Papina and R. van Woesik, 2003. Simultaneous time resolution of the emission spectra of fluorescent proteins and zooxanthellar chlorophyll in reef-building corals. Photochem. Photobiol., 77(5): 515-523.
- Gorbunov, M. and P. Falkowski, 2002. Photoreceptors in the cnidarian hosts allow symbiotic corals to sense blue moonlight. Limnol. Oceanogr., 47(1): 309-315.
- Gross, L., G. Baird, R. Hoffman, K. Baldridge and R. Tsien, 2000. The structure of the chromophore with DsRed, a red fluorescent protein from coral. Proc. Natl. Acad. Sci. USA, 97(22): 11990-11995.
- Gulko, D., M. Lesser and M. Ondrusek, 1995. Introduction of materials and methods commonly used by participants in the 1994 HIMB summer program on UV radiation and coral reefs. . In: Ultraviolet Radiation and Coral Reefs. D. Gulko and P.L. Jokiel (eds.), HIMB Technical Report #41, 19-23.
- Gurskaya, N., A. Fradkov, A. Terskikh, M. Matz, Y. Labas, V. Martynov, Y. Yanushevich, K. Lukyanov and S. Lukyanov, 2001. GFP-like chromoproteins as a source of far-red fluorescent proteins. FEBS Letters, 507(1):16-20.
- Gurskaya, N., V. Verkhusha, A. Shcheglov, D. Staroverov, T. Chepurnykh, A. Fradkov, S. Lukyanov and K. Lukyanov, 2006. Engineering of a monomeric green-to-red photoactivatable fluorescent protein induced by blue light. Nature Biotechnology, 24: 461-465.
- Heim, R., D. Prashner and R. Tsien, 1994. Wavelength mutations and posttranslational autoxidation of green fluorescent protein. Proc. Natl. Acad. Sci. USA, 91: 12501-12504.
- Henderson, J. and S. Remington, 2005. Crystal structures and mutational analysis of amFP486, a cyan fluorescent protein from Anemonia majano. Proc. Natl. Acad. Sci. USA, 102, 36: 12712-12717.
- Hochberg, E., M. Atkinson, A. Apprill and S. Andrèfouët, 2004. Spectral reflectance of coral. Coral Reefs, 23: 84-95.
- Hollingsworth, L., R. Kinzie III, T. Lewis, D. Krupp and J-AC Leong, 2005. Photoaxis of motile zooxanthellae to green light may facilitate symbiont capture by coral larvae. Coral Reefs, 24: 523.
- Ianushevich, I., N. Gurskaia, D. Staroverov, S. Lukyanov and K. Lukyanov, 2003. A natural fluorescent protein that changes its fluorescence during maturation. Bioorg Khim., 29(4):356-360.
- Ip, D., S-H Chan, M. Allen, M. Bycroft, D. Wan and K-B Wong, 2004. Crystallization and preliminary crystallographic analysis of a novel orange fluorescent protein from the cnidaria tube anemone Cerianthus. Acta Crystallogr., Section D, 60:340-341.
- Israel, A. and S. Beer, 1992. Photosynthetic carbon acquisition in the red alga Gracilaria conferta. 2. Rubisco carboxylase kinetics, carbonic anhydrase, and bicarbonate uptake. Mar. Biol., 112:697-700.
- Karasawa, S., T. Araki, M. Yamamoto-Hino, and A. Miyawaki, 2003. A green-emitting fluorescent protein from Galaxeidae coral and its monomeric use in fluorescent labeling. J. Biol. Chem., 278:34167-34171.
- Karasawa, S., T. Araki, T. Nagai, H. Mizuno and A. Miyawaki, 2004. Cyan-emitting and orange-emitting fluorescent proteins as a donor/acceptor pair for fluorescent resonance energy transfer. Biochem. J., 381(Part 1):307-312.
- Khang, S. and A. Salih, 2005. Localization of fluorescent pigments in a nonbioluminescent, azooxanthellate octocorals suggests a photoprotective function. Coral Reefs, 24, 3: 435.
- Kawaguti, S., 1937. On the physiology of reef corals II. The effect of light on colour and form of reef corals. Palao Trop. Biol. Sta. Taihoku Imperial University, Taihoku. 2.
- Kawaguti, S., 1944. On the physiology of reef corals VI. Study on the pigments. Palao Trop. Biol. Sta. Study, 2:617-674.
- Kawaguti, S., 1966. Electron microscopy on the fluorescent green of reef corals with a note on the mucous cells. Biol. J. Okayama Univ., 12:11-21.
- Kelmanson, I. and M. Matz, 2003. Molecular basis and evolutionary origins of color diversity in Great Star coral Montastraea cavernosa (Scleractinia:Faviida). Mol. Biol. Bull., 20, 7: 1125-1133.
- Kennedy, G.Y., 1979. Pigments of marine invertebrates. Adv. Mar. Biol., 16:309-381.
- Kinzie, R.A. and T. Hunter, 1987. Effect of light quality on photosynthesis of the reef coral Montiporaverrucosa. Mar. Biol., 94:95-109.
- Kinzie, R.A., P.L. Jokiel and R. York, 1984. Effects of light of altered spectral composition on coral zooxanthellae associations and on zooxanthellae in vitro. Mar. Biol., 78:239-248.
- Kirk, J.T.O., 1983. Light and Photosynthesis in Aquatic Ecosystems. Cambridge University Press, Cambridge. 401 pp.
- Koh, E., 1997. Secretion of bioactive compounds by a scleractinian coral. Proc. 8th Int. Coral Reef Symp., 2:1263-1266.
- Kutser, T., A. Dekker and W. Skirving, 2003. Modeling spectral discrimination of Great Barrier Reef benthic communities by remote sensing instruments. Limnol. Oceanogr., 48(1, Part 2): 497-510.
- Labas, Y., N. Gurskaya, Y. Yanushevich, A. Fradkov, K. Lukyanov, S. Lukyanov, and M. Matz, 2002. Diversity and evolution of the green fluorescent protein family. Proc. Natl. Acad. Sci. USA. 99 (7): 4256-4261.
- Larkum, A., G. Cox, R. Hiller, D. Parry and T. Dibbayawan, 1987. Filamentous cyanophytes containing phycocourobilin and in symbiosis with sponges and an ascidian of coral reefs. Mar. Biol., 95(1):1-13.
- Lesser, M., C. Mazel, D. Phinney and C. Yentsch, 2000. Light absorption and utilization by colonies of the congeneric hermatypic corals Montastraea faveolata and Montastraea cavernosa. Limnol. Oceanogr., 45(1): 76-86.
- Logan, A., K. Halcrow and T. Tomascik, 1990. UV excitation-fluorescence in polyp tissue of certain scleractinian corals from Barbados and Bermuda. Bull. Mar. Sci., 46(3):807-813.
- Louchard, E., R. Reid, C. Stephens, C. Davis, R. Leathers and T. Downes, 2003. Optical remote sensing of benthic habitats and bathymetry in coastal environments at Lee Stocking Island, Bahamas: A comparative spectral classification approach. Limnol. Oceanogr., 48(1):511-521.
- Lukyanov, K., A. Fradkov, N. Gurskaya, M. Matz, Y. Labas, A. Savitsky, M. Markelov, A. Zaraisky, X. Zhao, Y. Fang, W. Tan and S. Lukyanov, 2000. Natural animal coloration can be determined by a non-fluorescent green fluorescent protein homolog. J. Biol. Chem., 275(34):25879-25882.
- Manica, A. and R. Carter, 2000. Morphological and fluorescent analysis of the Montastraea annularis species complex in Florida. Mar. Biol., 137(5-6):899-906.
- Martynov, V., B. Maksimov, N. Martynova, A. Pakhomov, N. Gurskaya and S. Lukyanov, 2003. A purple-blue chromoprotein from Goniopora tenuidens belongs to the DsRed subfamily of GFP-like proteins. J. Biol. Chem., 278(47):46288-46292.
- Martynov, V., A. Savitsky, N. Martynov, P. Savitsky, K. Lukyanov and S. Lukyanov, 2001. Alternative cyclization in GFP-like proteins family. The formation and structure of the chromophore of a purple chromoprotein from Anemonia sculata. J. Biol. Chem., 276(24):21012-21016.
- Matz, M., A. Fradkov, Y. Labas, A. Savitsky, A. Zaraisky, M. Markelov and S. Lukyanov, 1999. Fluorescent proteins from nonbioluminescent Anthozoa species. Nature Biotechnology, 17:969-973.
- Matz, M., 2002. Diversity and evolution of the green fluorescent protein family. Proc. Natl. Acad. Sci. USA. 99 (7): 4256-4261.
- Matz, M., K. Lukyanov and S. Lukyanov, 2002. Family of the green fluorescent protein: Journey to the end of the rainbow. Bioessays 24(10):953-959.
- Matz, M., N. Marshall and M. Vorobyev, 2006. Are corals colorful? J. Am. Soc. Photochem. Photobiol., 82(2): 345-350.
- Mazel, C.H., 1995. Spectral measurements of fluorescence emission in Caribbean cnidarians. Mar. Ecol. Prog. Ser., 120:185-191.
- Mazel, C.H., 1997. Coral fluorescence characteristics: excitation – emission spectra, fluorescence efficiencies, and contribution to apparent reflectance. Ocean Optics XIII. 240-245.
- Mazel, C., M. Lesser, M. Gorbunov, T. Barry, J. Farrell, K. Wyman and P. Falkowski, 2003. Green fluorescent proteins in Caribbean corals. Limnol. Oceanogr., 48(1, part 2), 402-411.
- Mazel, C., M. Lesser, M. Gorbunov and P. Falkowski, 2004. Discovery of nitrogen-fixing cyanobacteria in corals. Science, 305: 997-1000.
- Mazel, C. H., and E. Fuchs, 2003. Contribution of fluorescence to the spectral signature and perceived color of corals. Limnol. Oceanogr. 48:390-401.
- Mazel, C.H., 2005. Underwater fluorescence photography in the presence of ambient light. Limnol. Oceanogr.: Methods 3:499-510.
- Minghelli-Roman, A., J. Chisolm, M. Marchioretti, and J. Jaubert, 2002. Discrimination of coral reflectance spectra in the Red Sea. Coral Reefs, 21(3):307-314.
- Miyawaki, A., 2002. Green fluorescent protein-like proteins in reef Anthozoa animals. Cell Struct. Func., 27:343-347.
- Moore, L. and S. Chisolm, 1999. Photophysiology of the marine cyanobacterium Prochlorococcus: Ecotypic differences among cultured isolates. Limnol. Oceanogr., 44(3):628-638.
- Moseley, H., 1877. On the colouring matter of various animals and especially of deep-sea forms dredged by the H.M.S. Challenger. Quarterly Journal of the Microscopical Society, 17: 1-23.
- Myers, M., J. Hardy, C. Mazel, and P. Dustan, 1999. Optical spectra and pigmentation of Caribbean reef corals and macroalgae. Coral Reefs 18: 2, 179-186.
- Nienhaus, K., B. Vallone, F. Renzi, J. Wiedenmann and G. Nienhaus, 2003. Crystallization and preliminary X-ray diffraction analysis of the red fluorescent protein eqFP611. Acta Cryst., D59:1253-1255.
- Nienhaus, K., G. Nienhaus, J. Wiedenmann and H. Nar, 2005. Structural basis for photo-induced protein cleavage and green-to-red conversion of fluorescent protein EosFP. Proc. Natl. Acad. Sci., USA, 102(26):9156-9159.
- Neveux, J., M. Tenório, C. Dupouy and T. Villareal, 2006. Spectral diversity of phycoerythrins and diazotroph abundance in tropical waters. Limnol. Oceanogr., 51(4):1689-1698.
- Pakahomov, A., N. Martynova, N. Gurskaya, T. Balashova and V. Martynov, 2004. Photoconversion of the chromophore of a fluorescent protein from Dendronephthya sp. Biochem. (Mosc.), 69(8):901-908.
- Papina, M., Y. Sakihama, C. Bena, R. van Woesik and H. Yamasaki, 2002. Separation of highly fluorescent proteins by SDS-PAGE in Acroporidae corals. In press, Comp. Biochem. Physiol.
- Paringit, E. and K. Nadaoka, 2001. Development of a canopy reflectance model for coral reef areas: Inferences from field spectral measurements and modeling efforts. Proc. 22nd Asian Conference on Remote Sensing.
- Pasternak, Z., B. Blasius, A. Abelson and Y. Achituv, 2006. Host-finding behavior and navigation capabilities of symbiotic zooxanthellae. Coral Reefs, 25(2):201-207.
- Pederson, H., 1968. Fluorescent flowers of the sea. Sea Frontiers, 14(4): 194-199.
- Peelle, B., T. Gururaja, D. Payan and D. Anderson, 2001. Characterization and use of green fluorescent proteins from Renilla mulleri and Ptilosarcus guernyi for the human cell display of functional peptides. J. Protein Chem., 20(6):507-19.
- Pieribone, V. and D. Gruber, 2005. Aglow in the Dark: The Revolutionary Science of Biofluorescence. The Belknap Press of Harvard University Press, Cambridge Massachusetts, and London, England. 263 pp.
- Piniak, G., N. Fogarty, C. Addison and W. Kenworthy, 2005. Fluorescence census techniques for coral recruits. Coral Reefs, 24: 496-500.
- Prashner, D., V. Enckenrode, W. Ward, F. Pentegast and M. Cormier, 1992. Primary structure of the Aequorea victoria green-fluorescent protein. Gene, 111: 229-233.
- Riddle, D. and A. Amussen, 1998. Coral pigments. Online presentation.http://www.reefs.org/library/talklog/d_riddle_042698.html
- Riddle, D., 2003. Effects of narrow bandwidth light sources on coral host and zooxanthellae pigments. Advanced Aquarist Online, November 2003. http://www.advancedaquarist.com/issues/nov2003/feature.htm
- Riddle, D., 2006. Product Review: Ocean Optics spectrometers and software. Advanced Aquarist Online, June 2006. http://www.advancedaquarist.com/2006/6/review
- Rinkevich, B. and Y. Loya, 1983. Intraspecific competition networks in the Red Sea coral Stylophora pistillata. Coral Reefs, 1:161-172.
- Salih, A., 2003. An exploration of light regulating pigments of reef-building corals from macro- to micro- and nano-scales. In: From Zero to Infinity, J. Nicholls and B. Pailthorpe, eds. The Science Foundation for Physics, University of Sydney. Chapter 4:49-69.
- Salih, A., A. Larkum, T. Cronin, J. Wiedenmann, R. Szymczak and G. Cox, 2004. Biological properties of coral GFP-type proteins provide clues for engineering novel optical probes and biosensors. In: Genetically Engineered and Optical Probes for Biomedical Applications II, A. Savitsky et al., eds. Proc. of SPIE, 5329:61-72.
- Salih, A., O. Hoegh-Guldberg and G. Cox, 1998. Photoprotection of symbiotic dinoflagellates by fluorescent pigments in reef corals. Proc. Australian Coral Reef Society, 75th Anniversary Conference, Heron Island, Australia. 217-230.
- Salih, A., A. Larkum, G.Cox, M.Kuhl and O.Hoegh-Guldberg, 2000. Fluorescent pigments in corals are photoprotective. Nature, 408: 850-853.
- Salih A, Larkum AWD & G Cox, 2001. Photoprotection from photoinhibition of symbiotic algae in corals by fluorescent pigments. 12th International Congress on Photosynthesis, Brisbane, Australia.
- Salih, A., G. Cox and A. Larkum, 2003. Cellular organization and spectral diversity of GFP-like proteins in live coral cells studied by single and multiphoton imaging and microspectroscopy. In: Multiphoton Microscopy in the Biomedical Sciences III, A. Periasamy and P. So, Editors. Proc. SPIE, Vol. 4963: 194-200.
- Salih, A, Larkum AWD & G Cox, 2001. Corals use fluorescent pigments as natural sunscreens. Australian Coral Reef Society Conference, Magnetic Island, Australia.
- Salih, A, Cox G & AWD Larkum, 2000. The role of fluorescent pigments: evidence of enhanced resistance to bleaching in fluorescently pigmented corals. IX International Coral Reef Symposium, Bali, Indonesia.
- Salih, A., G. Cox & AWD Larkum, 2000. Energy dissipation by fluorescence coupling by coral fluorescent pigments. Optical Society Conference, Application of Optical Techniques in Biological Sciences.
- Salih, A., Hoegh-Guldberg O, Cox G & AWD Larkum, 1999. Protection against bleaching of corals by fluorescent pigments during the 1998 mass bleaching event. XIX Pacific Science Congress.
- Salih A., Hoegh-Guldberg, O. & Cox G., 1999. Are fluorescent colors in corals photoprotective? Australian Coral Reef Society Conference.
- Salih, A., Hoegh-Guldberg, O. & Cox G., 1998. Do fluorescent pigments in corals provide photoprotection to their algal endosymbionts and reduce the severity of bleaching? Australian Coral Reef Society Conference.
- Salih, A., Hoegh-Guldberg O & Cox G., 1997. Bleaching responses of corals: the effects of light and elevated temperature on their morphology and physiology. Australian Coral Reef Society Conference.
- Salih, A., Hoegh-Guldberg O & Cox G., 1997. Photoprotection of symbionts by fluorescent sunscreens in reef corals. Australian Coral Reef Society 75th Anniversary Conference.
- Schlichter, D., W. Weber and H.W. Fricke, 1985. A chromatophore system in the hermatypic, deep water coralLeptoseris fragilis (Anthozoa: Hexacorallia). Mar. Biol. 89:143-147.
- Schlichter, D., H.W. Fricke and W. Weber, 1986. Light harvesting by wavelength transformation in a symbiotic coral of the Red Sea twilight zone. Mar. Biol., 91:403-407.
- Schonwald, H., Y. Achituv, and Z. Dubinsky, 1987. Differences in the symbiotic interrelation in dark and light coloured colonies of the hydrocoral Millepora dichotoma. Symbiosis 4:171-184.
- Shagin, D., E. Barsova, Y. Yanushevich, A. Fradkov, K. Lukyanov, Y. Labas, T. Semenova, J. Ugalde, A. Meyers, J. Nunez, E. Widder, S. Lukyanov and M. Matz, 2004. GFP-like proteins as ubiquitous metazoan superfamily: Evolution of functional features and structural complexity. Mol. Biol. Evol., 21(5):841-850.
- Shibata, K., 1969. Pigments and a UV-absorbing substance in corals and a blue-green alga living in the Great Barrier Reef. Plant and Cell Physiol., 10: 325-335.
- Shimomura, O., F. Johnson and Y. Saiga, 1962. J. Cell Comp. Physiol. 59:223-239.
- Shkrob, M., Y. Yanushevich, D. Chudakov, N. Gurskaya, Y. Labas, S. Poponov, N. Mudrik, S. Lukyanov and K. Lukyanov, 2005. Far-red fluorescent proteins evolved from a blue chromoprotein from Actinia equina.Biochem. J., On-line publication.
- Sole-Cava, L.M., and J.P. Thorpe, 1992. Genetic divergence between the color morphs in populations of the common intertidal sea anemones Actinia equina and A. prasina (Anthozoa: Actiniaria) in the Isle of Man. Mar. Biol. 112:243-252.
- Sprung, J. and J. Delbeek, 1997. The Reef Aquarium – A Comprehensive Guide to the Identification and Care of Tropical Marine Invertebrates, Vol. 2. Ricordea Publishing, Coconut Grove, Florida. 546 pp.
- Takabayashi, M. and O. Hoegh-Guldberg, 1995. Ecological and physiological differences between the two colour morphs of the coral Pocillopora damicornis. Mar. Biol., 123: 705-714.
- Takabayashi, M., D. Carter, J. Lopez and O. Hoegh-Guldberg, 2003. Genetic variation of the scleractinian coralStylophora pistillata, from western Pacific reefs. Coral Reefs 22(1): 17-22.
- Terskikh, A., A. Fradkov, A. Zaraisky, A. Kajava and B. Angres, 2002. Analysis of red mutants. Space around the fluorophore accelerates fluorescence development. J. Biol. Chem., 277(10):7633-7636.
- Todd, P., R. Sidle and L. Chou, 2002. Plastic corals from Singapore: 1. Coral Reefs, 21(4):391.
- Todd, P., R. Sidle and L. Chou, 2002a. Plastic corals from Singapore: 2. Coral Reefs, 21(4):407.
- Todd, P., R. Sidle and L. Chou, 2003. Erratum: Plastic corals from Singapore: 1. Coral Reefs, 22(3):306.
- Tsien, R., 1998. The green fluorescent protein. Annu. Rev. Biochem., 67:509-544.
- Tsutsui, H., S. Karasawa, H. Shimizu, N. Nukina and A. Miyawaki, 2005. Semi-rational engineering of a coral fluorescent protein into an efficient highlighter. EMBO Reports. 6:233-238.
- Tu, H., Q. Xiong, S. Zhen, X. Zhong, L. Peng, H. Chen, X. Jiang, W. Liu, W. Yang, J. Wei, M. Dong, W. Wu and A. Xu, 2003. A naturally enhanced green fluorescent protein from magnificent sea anemone (Heteractis magnifica) and its functional analysis. Biochem. Biophys. Res. Commun., 301(4):879-885.
- Tyree, S., 1998. Reef Building Stony Corals: The Natural Physical Environment. A Review of the Research Papers. D.E. Publishing, Murrieta, Ca. 418 pp.
- Verkhusha, V. and K. Lukyanov, 2004. The molecular properties and applications of Anthozoa fluorescent proteins and chromoproteins. Nature Biotechnology, 22(3):289-296.
- Vermeij, M., L. Delvoye, G. Nieuwland and R. Bak, 2002. Patterns in fluorescence over a Caribbean reef slope: The coral genus Madracis. Photosynthetica, 40(3): 423-429.
- Veron, J., 1993. Corals of Australia and the Indo-Pacific. University of Hawaii Press, Honolulu. 644 pp.
- Veron, J., 1995. Corals In Space and Time – The Biogeography and Evolution of the Scleractinia. Cornell University Press, Ithaca, NY. 321 pp.
- Vestheim, H. and S. Kaartvedt, 2006. Plasticity in coloration as an antipredator strategy among zooplankton. Limnol. Oceanogr., 5(14): 1931-1934.
- Wachter, R. and S. Remington, 1999. Sensitivity of the yellow variant of GFP to halides and nitrate. Curr. Biol., 9(17); R627-R630.
- Weber, W. and H. Gras, 1980. Ultrastructure observations on changes in cell shapes in chromatophores of the Sea Urchin Centrostephanus longispinus. Cell Tiss. Res., 206:21-33.
- Weber, W. and M. Daumbach, 1974. Light-sensitivity of isolated pigment cells of the Sea UrchinCentrostephanus longispinus. Cell Tiss. Res., 148:437-440.
- Wiedenmann, J., C. Elke, K-D. Spindler, and W. Funke, 2000. Cracks in the beta-can: Fluorescent proteins from Anemonia sulcata (Anthozoa, Actinaria). Proc. Natl. Acad. Sci. USA, 97(26): 14091-14096.
- Wiedenmann, J., A. Schenk, C. Röcker, A. Girod, K-D. Spindler and G. Nienhaus, 2002. A far-red fluorescent protein with fast maturation and reduced oligomerization tendency from Entacmaea quadricolor(Anthozoa, Actinaria). Proc. Natl. Acad. Sci. USA, 99, 18: 11646-11651.
- Wiedenmann, J., A. Schenk, C. Röcker, A. Girod, K-D. Spindler and G. Nienhaus, 2002a. Correction for: A far-red fluorescent protein with fast maturation and reduced oligomerization tendency from Entacmaea quadricolor (Anthozoa, Actinaria). Proc. Natl. Acad. Sci. USA, 99, 20:14091-14096.
- Wiedenmann, J., S. Ivanchenko, F. Oswald, F. Schmitt, C. Röcker, A. Salih, K-D. Splinder and G. Nienhaus, 2004. EosFP, a fluorescent marker protein with UV-inducible green-to-red fluorescence conversion. Proc. Natl. Acad. Sci. USA, 101(45):15905-15910.
- Wilmann, P., J. Battad, T. Beddoe, S. Olsen, S. Smith, S. Dove, R. Devenish, J. Rossjohn and M. Prescott, 2005. The 2.0 Å crystal structure of a pocilloporan at pH 3.5: The structural basis for the linkage between color transition and halide binding. Photochem. Photobiol., 82(2):359-66.
- Yampolsky, I., S. Remington, V. Martynov, V. Potapov, S. Lukyanov and K. Lukyanov, 2005. Synthesis and properties of the chromophore of the asFP595 chromoprotein from Anemonia sulcata. Biochem., 44(15):5788-5793.
- Yanushevic, Y., M. Bulina, N. Gurskaya, A. Savitskii and K. Lukyanov, 2002. Key amino acid residues responsible for the color of green and yellow fluorescent proteins from the coral polyp Zoanthus sp. Russ. J. Bioorg. Chem., 28(4): 274-277.
- Yentsch, C. and D. Phinney. CoBOP Coral Reefs: Optical closure of a coral reef submarine light field. On-line publication.
- Yu, J-K, T-H Liao, C. Chen, T-H Liao, L-S Fang and W-S Tsai, 2000. Do color patterns of Pocillopora damicornisreflect zooxanthellae diversity? Coral Reefs, 19(1): 98-99.
- Yu, M-H., A. Glazer, K. Spencer and J. West, 1981. Phycoerythrins of the red alga Callithamnion. Plant Physiol., 68:482-488.
- Zawada, D. and J. Jaffe, 2003. Changes in the fluorescence of the Caribbean coral Montastraea faveolataduring heat-induced bleaching. Limnol. Oceanogr., 481(1, part 2):412-425.
- Zimmer, M., 2005. Glowing Genes: A Revolution in Biotechnology. Prometheus Books, Amherst, New York. 222 pp.




0 Comments